Detection and quantification of extracellular microRNAs in medulloblastoma
Abstract
Aim: Medulloblastoma (MB) is the most common malignant brain tumor in children. The crucial role of extracellular-microRNAs (ex-miRNAs) in cancer has been widely recognized; however, their role in MB remains unknown. This study aimed to investigate MB-driven ex-miRNAs.
Methods: Microarray analysis was used to disclose the identity and quantity of key miRNAs excreted in culture-medium (CM) of 3 human MB cell lines and cerebrospinal fluid (CSF) of brain tumors (including MB) and leukemia patients. MiRNA expression was validated by quantitative reverse transcription polymerase chain reaction.
Results: We have demonstrated that the 3 MB cell lines tested commonly expressed 1,083 miRNAs in their spent CM. Among them, 57 miRNAs were specific to the CM of metastasis-related cell lines which represents the aggressive group 3 and group 4 MB subtypes. A significant number (1,254) of ex-miRNAs were identified in the CSF of a MB patient. Eighty-six of these miRNAs were found to be differentially expressed in this patient’s CSF compared with controls. Interestingly, 3 metastasis-associated miRNAs over-represented in CM of metastasis-related MB cell lines were found to be significantly enriched in the CSF of the MB patient.
Conclusion: Although more samples are required to fully verify these results, our work provides the first evidence for the presence of a significant amount of miRNAs excreted extracellularly by MB cells and raises the possibility that, in the near future, miRNAs could be probed in CSF of MB patients and serve as novel biological markers.
Keywords
Introduction
Medulloblastoma (MB) is the most common malignant brain tumor in children.[1] Metastatic MB carries a poor prognosis.[2] Mechanisms that predict dissemination are poorly understood. Recently, several studies have revealed a critical role for microRNAs (miRNAs) during tumorigenesis and metastasis of several cancers, including MB.[3-6]
Besides intracellular miRNAs with the traditional function of translation regulation, there is accumulating evidence that miRNAs exist extracellularly in body fluids, including cerebrospinal fluids (CSF).[7,8] Several reports have described that deregulated extracellular-miRNAs (ex-miRNAs) are closely associated with the clinical course of malignant tumors.[9,10] Interestingly, such deregulation returns to a normal level after tumor resection.[7,8] Hence, expression analysis of ex-miRNAs is of increasing interest for diagnostic and prognostic purposes.
Every cancer investigated has a distinct miRNA signature and deregulated levels of miRNAs have been detected in body fluids of patients, including those with lymphoma,[11] leukemia,[12] colon,[13] breast,[14] prostate,[15] ovarian,[16] pancreatic,[17] gastric,[18] and lung cancer.[19] In the context of brain tumors, recent studies have demonstrated a significant presence of certain miRNAs in CSF samples from patients with central nervous system lymphoma, glioma, and metastatic brain cancers.[20-22] Recent miRNA profiling of CSF has enabled early detection of glioblastoma and reflected disease activity.[22] Therefore, ex-miRNAs may represent important minimally invasive candidate biomarkers in brain tumors. The presence and biological role of ex-miRNAs in MBs, however, remain unknown. This study was conducted to gain insight into the identity and quantity of MB-related ex-miRNAs and to speculate on their possible biological function in the context of MB metastasis.
Methods
Patient characteristics and CSF
CSF samples from patients with MB (n = 2), control patients with leukemia with no intracranial mass lesions and/or neurologic disorders (n = 3), CSF samples from patients with ependymoma (n = 3) and glioblastoma (n = 1) that were collected from patients treated at the University Children’s Hospital of Zürich, Switzerland. Written informed consent was obtained from each patient. CSF samples from patients with MB were collected 3 weeks after surgery and before start of radiotherapy or chemotherapy. CSF samples were centrifuged (500 g, 10 min, room temperature) within 60 min after collection to remove cells and debris and were stored at -80 °C until further processing.
Human MB cell lines
Human MB cell lines (DAOY and D283) were purchased from American Type Culture Collection (Manassas, VA, USA). D341 human MB cells were the kind gift of Dr. Henry Friedman (Duke University, Durham, UK). MB cell lines were cultured as previously published[23] and maintained at 37 °C in a humidified atmosphere with 5% CO2. To isolate RNA from cultured medium, 10.000-20.000/mL DAOY cells or 20.000-40.000/mL D341, D283, and T293 cells were plated and left to grow in their conditioned media for 72 h in 24 wells plates. Conditioned medium (2 mL) of each cell lines were centrifuged at 1,200 rpm to remove cells. The supernatant was then centrifuged at 10,000 rpm to remove debris.
RNA extraction for microarray
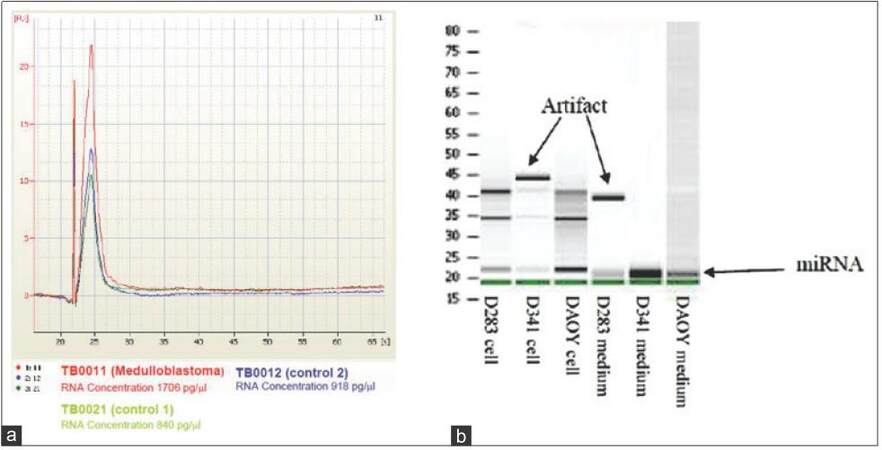
Total RNA from cell cultures or CSF were extracted using a mix of Qiazol, Qiagen (Qiagen, Basel, Switzerland) and chloroform directly on cells. For small RNA in conditioned medium or CSF, the addition of miRNAs extraction reagent (Toray) was performed. In both situations, a centrifugation step was required to collect aqueous phase containing RNA that was finally transferred to miRNeasy Mini spin column from miRNeasy purification kit Qiagen (Qiagen, Basel, Switzerland). After subsequent washing steps, RNAs were eluted using 30 μL of nuclease-free water and concentrated up to 3 μL with vacuum concentrator. Quality was checked on Bioanalyzer using RNA 6000 Pico Chip (Agilent Chemical Analysis, Life Sciences, and Diagnostics. Basel, Switzerland) gel and quantified using Nanodrop Photometer [Figure 1a and b].
Labeling and hybridization
Total RNA (250 ng) extracted from cells and 3 μL of concentrated small RNA extracted from medium were used with Toray 3D-Gene miRNA labeling kit (Toray, Japan) in presence of spikes used as positive controls. Briefly, 5’-phosphates were removed from miRNA end using alkaline phosphatase and a fluorescent label was enzymatically attached to the 3’-end of the miRNA. After an enzyme inactivation step and addition of a hybridization buffer, labeled miRNA was injected on 3D-Gene Human miRNA Oligo Chips (Toray, Japan) targeting 2019 miRNA based on miRBase release 19. Finally, arrays were placed in a hybridization chamber and set into a 32 °C oven for 16 h with a shaker adjusted to 250 rpm.
Washing and scanning
Arrays were washed using 3 solutions with different stringencies to remove non-specifically bound miRNAs. Then, arrays were scanned with the 3D-Gene Scanner 3000 instrument (Toray, Japan) to measure fluorescence. Scanning was carried out using 3 different photomultiplier sensitivities (PMT gain) to allow optimizing of signal detection and checking for consistency.
Microarray analysis
Images were analyzed with the 3D-Gene Extraction software (Toray, Japan). After completion of the auto-analysis work followed on image files, raw signals, and detection calls was produced in tabular files. GeneSpring GX12 (Agilent) was then used to apply quantile normalization and differential expression analysis using modified t-test implemented in the software. Experimental variability was assessed with principal component analysis (PCA) [Figures 2 and 3b] and Pearson correlation matrix [Figure 3a] generated using the same software.
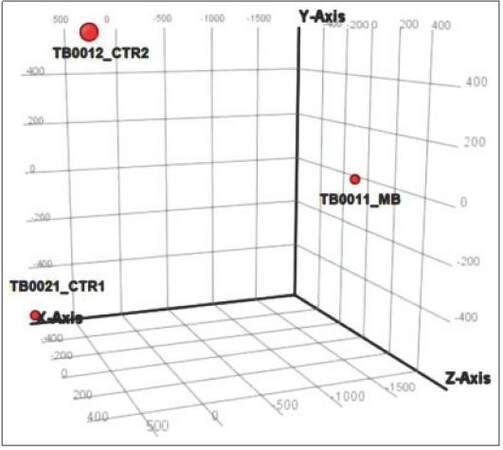
Figure 2. PCA graph showing microRNA spectra in CSF of MB patient vs. control CSFs. TB0021_CTR1: CSF from patient with no brain tumor control 1; TB0012_CTR2: CSF from patient with no brain tumor control 2; TB0011_ MB: CSF of MB patient. MB: Medulloblastoma; CSF: cerebral spinal fluid; PCA: Principal component analysis
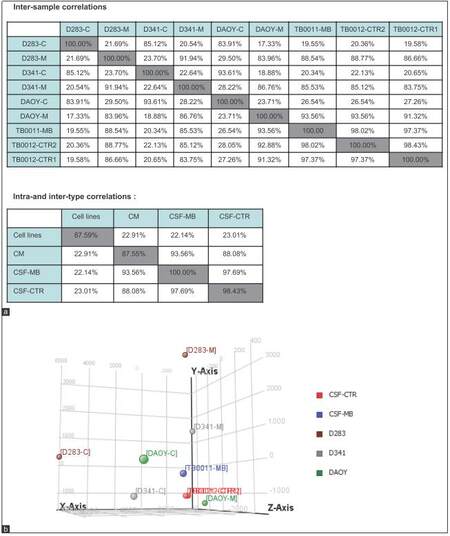
Figure 3. (a) Pearson correlation analysis for indicated samples; (b) PCA graph representing microRNA spectra inside MB cell lines (cell line name-C), in culture medium (cell line name-M) and in CSF of MB patient (TB0011-MB) compared to control TB0021_CTR1 and TB0012_CTR2. Graph demonstrating similarity between miRNA profile in CM and those secreted in CSF of MB patient. CM: Culture medium; PCA: Principal component analysis; MB: Medulloblastoma; CSF: Cerebral spinal fluid
MicroRNA isolation for reverse transcription polymerase chain reaction analysis
For precipitation of nucleic acids, the monovalent cation concentration of the solution was adjusted to 0.5 mol/L sodium acetate. Glycogen (AM9510, Ambion, Life Technology, NY, USA) was added to a final concentration of 100 μg/mL. The solution was then mixed with 1 volume of isopropanol. The mixture was chilled for 20 min at -20 °C, then centrifuged for 20 min at 13,000 rpm. The supernatant fluid was removed, and the nucleic acid resuspended in lysis buffer. Final purification of RNA enriched for small RNAs from 600 μL of conditioned media and CSF samples was obtained using the mirVanaTM miRNA Isolation Kit (Ambion, Life Technology) according to manufacturer’s instructions for “Enrichment Procedure for Small RNAs.” Using this approach consisting of two sequential filtrations with different ethanol concentrations, an RNA fraction highly enriched in RNA species ≤ 200 nt was obtained. First strand synthesis of mature miRNAs was followed by quantitative reverse transcription polymerase chain reaction (qRT-PCR) using miRNA-specific TaqMan MGB probes (Applied Biosystems, Life Technology). For the qRT-PCR reaction, the Gene Expression Master Mix was used and the protocol was optimized for the ABI7900HT reader (Applied Biosystems). Probe-primer solutions specific for the following miRNAs were used: miR-1290 (002863), miR-125a- 3p (002199), miR-1298 (002861), miR-125b-1* (002378), miR-486-3p (002093), miR-572 (001614), miR-4476 (464702_mat), miR-615-5p (002353), and miR-3918 (464506_mat) (Applied Biosystems, Life Technology). The relative gene expression was calculated for each gene of interest using the ΔΔCT method, where cycle threshold values were normalized to the level of cel-miR-39-3p (4464066, Ambion, Life Technology), which was used as spike-in by adding it during the lysis step of miRNAs extraction.
Results
Detection of ex-miRNAs in cultured medium of MB cell lines by microarray analysis
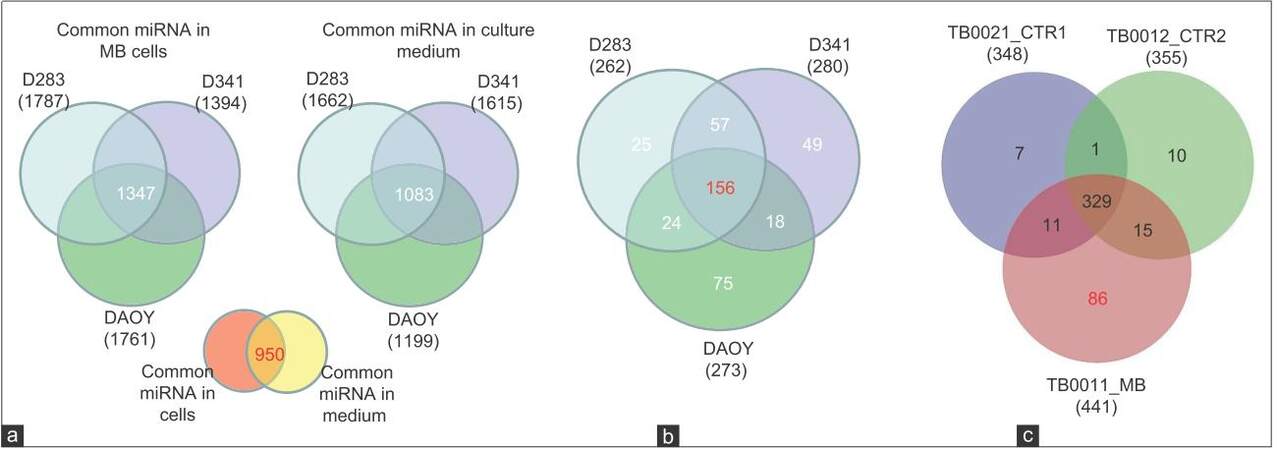
Given that some human cancer cells secrete miRNAs into their extracellular environment and body fluids,[24-26] it was hypothesized that MB cell lines may secrete miRNAs into their spent culture medium. To test this hypothesis, 3 cell lines representing MB subtypes D341 and D283 (metastasis-related group 3 and group 4 MB subtypes)[27] and DAOY (sonic hedgehog-related) were ultured individually for 72 h in vitro and miRNAs expression was analyzed in the lysates of each MB cell line and in their corresponding culture media. We identified 1,662, 1,615, and 1,199 secreted miRNAs in the culture-medium (CM) of MB cell lines D283, D341 and DAOY, respectively, among them 1,083 miRNAs that were common in the CM of the 3 cell lines. In cell lysates of D283, D341 and DAOY, on the other hand, we detected 1,787, 1,394 and 1,761 miRNA respectively, with 1,347 miRNAs found common to all 3 cell lines [Figure 4a]. Interestingly, 950 miRNAs were commonly identified in CM of both groups and in lysates of the 3 cell lines tested, indicating that the level of ex-miRNAs may well reflect the expression level of tumor miRNAs. Using a fold-change > 2, we identified a group of 156 miRNAs that are commonly enriched in CM derived from the 3 cell lines compared to their respective cell lysates [Figure 4b] and [Supplementary Table 1] and 57 miRNAs that were specific to the CM of D341 and D283, which represented the 2 metastasis-related group 3 and group 4 MB subtypes, respectively[27] compared to DAOY-derived CM [Figure 4b] and [Supplementary Table 2]. We found 2 additional groups of miRNAs to be differentially enriched in CM of D341 and D283, represented by 60 miRNAs overrepresented and 52 underrepresented compared to DAOY-derived CM [Supplementary Tables 3 and 4]. Overall, the results of this experiment demonstrate that MB cell lines secrete miRNAs into the CM and that certain ex-miRNAs retain different enrichment levels in the CM-derived from the 2 cell lines representing the metastasis-related group 3 and group 4 MB subtypes.
Figure 4. MiRNAs expression spectrum in MB cell lines, their corresponding CM and in CSF (a) Quantitative Venn diagram showing miRNAs commonly expressed in indicated cell lines and their corresponding CM. Differences linked to the expression level rather than detection threshold; (b) Venn diagram showing that 156 miRNAs enriched in CM of MB cell lines compared to their cell lysates and 57 miRNAs enriched in CM of the MR D283 and D341 but not in DAOY. Fold change > 2; (c) Venn diagram demonstrating miRNAs with high expression in CSF of MB patient compared to controls. TB0021_CTR1: CSF of leukemia with no brain tumor (patient control 1); TB0012_CTR2: leukemia with no brain tumor (patient control 2); TB0011_MB: CSF of MB patient. MB: Medulloblastoma; CSF: Cerebral spinal fluid; CM: Culture-medium; MiRNAs: MicroRNAs
Detection of ex-miRNAs in CSF of MB patients by microarray analysis
We next asked whether ex-miRNAs could be detected in CSF of MB patients, to test whether it would be technically possible to use the CSF as a source for diagnostic miRNA testing. Using microarray analysis, we screened cell-free CSF from a patient with MB and compared the results to controls (CSF from two different leukemia patients with no cerebral manifestation or neurological disease). PCA [Figure 2] showed clear separation between the miRNA spectrum in CSF of MB patient and controls. Microarray analysis identified 1,254 miRNAs in the MB-CSF sample [Table 1], of which 86 miRNAs were differentially expressed in CSF of the MB patient compared to the 2 CSF controls [Figure 4c] and [Supplementary Table 5]. Further analysis identified 268 miRNAs over-represented (with fold-change > 2) and 6 miRNAs under-represented in MB-CSF compared with the 2 different controls tested [Supplementary Tables 6 and 7], indicating a trend toward miRNA enrichment in the MB-CSF sample.
Number of miRNAs detected in CSF
| CSF | Number of miRNA detected |
|---|---|
| CSF of MB patient | 1,254 |
| CSF control 1 | 1,004 |
| CSF control 2 | 1,049 |
Comparison between miRNA expressions in CM vs. CSF samples
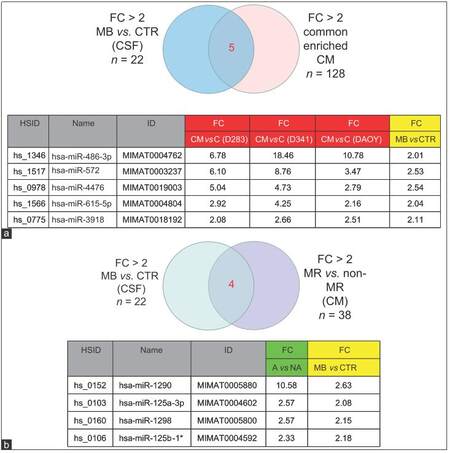
An overlap of the spectra of ex-miRNA candidates detected in the CSF of the MB patient and those excreted by MB cell lines into the CM would support our hypothesis of miRNA secretion by MB cells. Indeed, Pearson correlations analysis showed that ex-miRNAs profiles in MB-CSF displayed a good homogeneity with the profile of miRNAs secreted in CM of MB cell lines [Figure 3a]. This conclusion was confirmed by PCA showing clear separation of miRNAs derived from lysates of MB cell lines from those derived of MB-CSF samples or of CM derived from MB cells [Figure 3b], confirming the conclusions from Pearson correlations. Compiling expression tables allowed identification of 5 miRNAs (miR-486-3p, miR-572, miR-3918, miR-4476, and miR-615) that were significantly up-regulated in the CM of the 3 cell lines (D283, D341 and DAOY) and enriched in MB-CSF compared to control CSF [Figure 5a]. Moreover, 3 other miRNAs (miR-1290, miR-125a, miR-125b), known to be associated with metastasis, and miR-1298, were over-represented in the CM of metastasis-related cell lines (D283 and D341), but not in DAOY and were significantly over-represented in MB-CSF compared to control CSF [Figure 5b].
Figure 5. MiRNAs commonly enriched in CM of MB cell lines, and in CSF sample. (a) Venn diagram and table presenting 5 miRNAs commonly upregulated in CM of 3 MB cell lines and in CSF from MB patients compared to CTR; (b) Venn diagram and table representing 4 miRNAs commonly upregulated in CSF samples from MB patients and over-represented in CM of the MR cell lines D341, D283. CM: Culture-medium; MB: Medulloblastoma; CSF: Cerebral spinal fluid; MR: Metastasis related; FC: Fold change; CTR: Control; MiRNAs: MicroRNA
Validation of microarray data by qRT-PCR
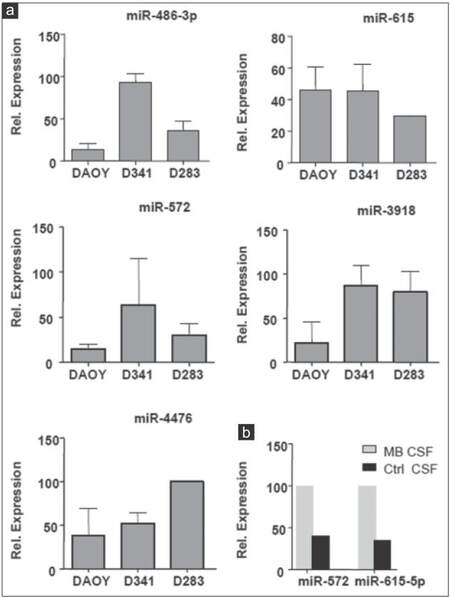
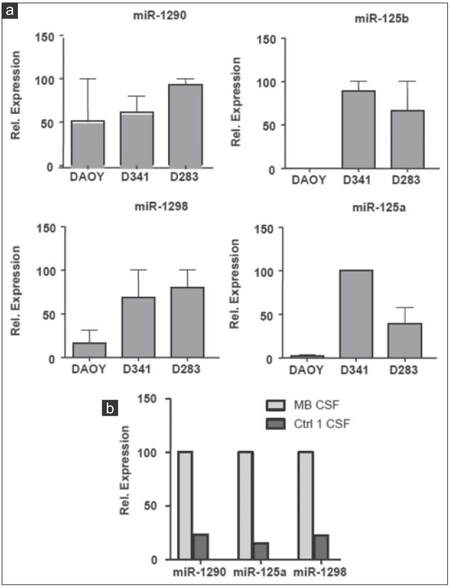
In order to further verify the results of miRNA microarray analysis, we selected miR-486-3p, miR-572, miR-3918, miR-4476, and miR-615 for quantitative real-time PCR assays because of their over-representation in the CM of the 3 MB cell lines and in MB-CSF. TaqMan analysis confirmed the outcomes of miRNA microarray profiling for the 5 miRNAs tested and showed that the five were enriched in the CM of the 3 cell lines [Figure 6a]. However, only miR-615 and miR-572 were also accumulated in MB-CSF [Figure 6b]. We also chose miR-1290, miR-125a, miR-125b, and miR-1298 as other candidates for qRT-PCR due to their over-representation in CM of metastasis-related cell lines D283 and D341, as well as in CSF of the MB patients. TaqMan analysis showed an evidently increased expression level of the 4 cell line-derived miRNAs in both D283 and D341 compared to DAOY [Figure 7a]. The levels of 3 miRNAs (miR-1290, miR-125a, miR-1298) were also markedly increased in MB-CSF, thus confirming the results of the miRNA microarray analysis for 3 out of 4 of these selected miRNAs [Figure 7b].
Figure 6. Expression analysis of 5 ex-miRNAs in MB cell lines CM and in CSF of MB patient. (a) TaqMan qRT-PCR analysis for miR-486-3p, miR-572, miR-3918, miR-4476, miR-615 in CM of indicated cell lines; (b) TaqMan qRT-PCR analysis for miR-572, miR-615 in MB-CSF (n = 3; ± standard deviation). MB: Medulloblastoma; CSF: Cerebral spinal fluid; CM: Culture-medium; qRT-PCR: Quantitative reverse transcription polymerase chain reaction
Figure 7. Expression analysis of 4 overexpressed ex-miRNAs in MB cell lines CM and in CSF of MB patient. (a) TaqMan qRT-PCR analysis for miR-1290, miR-125a, miR-1298, miR-125b in CM of indicated cell lines; (b) TaqMan qRT-PCR analysis for miR-1290, miR-125a, miR-1298 in MB-CSF (n = 3; ± standard deviation). MB: Medulloblastoma; CSF: Cerebral spinal fluid; CM: Culture-medium; qRT-PCR: Quantitative reverse transcription polymerase chain reaction
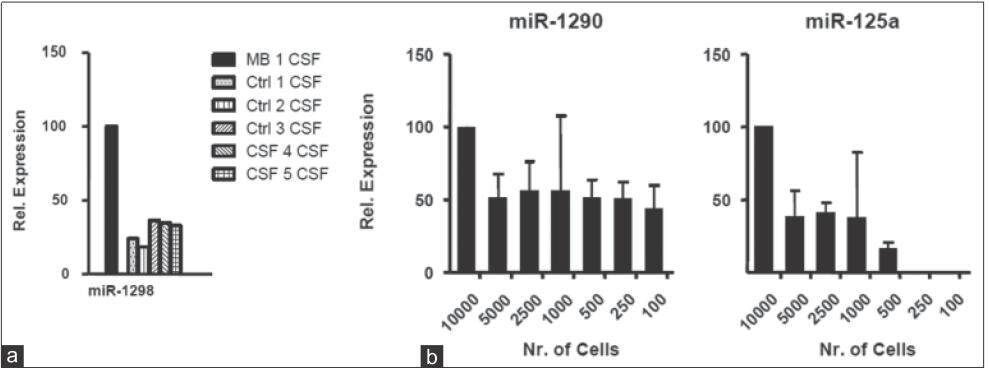
To further validate the finding of selective enrichment of miR-1298 in MB-CSF, we tested it against an additional 5 different CSF controls from one leukemia patient, 3 ependymoma patients, and one glioblastoma patient (specifically chosen to control for brain surgery as a possible factor influencing miRNA secretion). Consistently, TaqMan analysis confirmed significant enrichment of miR-1298 in MB-CSF compared to the 5 controls [Figure 8a]. Together, using TaqMan analysis, we confirmed the microarray data result and demonstrated the feasibility of quantitative detection of miRNAs in culture medium and CSF using qRT-PCR (popular gene expression assay and efficient method for high-throughput used in most diagnostic labs). Importantly, we could detect ex-miRNAs by qRT-PCR in CM of as few as 100-500 MB cells [Figure 8b], recommending qRT-PCR for the development of non-invasive detection of metastasis-predicting markers for MB.
Figure 8. Relative ex-microRNAs expression analysis in CSF, MB cell lines: (a) TaqMan qRT-PCR analysis for miR-1298 compared to 5 controls: Ctrl 1 CSF from leukemia patient, Ctrl 2 CSF from glioblastoma patient, Ctrl 3-5 CSF from 3 ependymoma patients (n = 3; ± SD); (b) TaqMan qRT-PCR analysis for miR-1290, miR-125a in serial dilution of D341 CM (n = 3; ± SD). CM: Culture-medium; MB: Medulloblastoma; CSF: Cerebral spinal fluid; qRT-PCR: Quantitative reverse transcription polymerase chain reaction; SD: standard deviation
Discussion
Aberrant expression of ex-miRNA circulating in CSF of certain brain tumor patients has recently been reported to be cancer biomarkers and potential regulators of the disease.[7,8] However, the existence and role of ex-miRNAs in MB extracellular environment are unknown. Therefore, better understanding of ex-miRNA secretion and function in MB seems crucial for the development of novel insights for its diagnosis and prognosis. This study aimed to identify key miRNAs in culture medium of 3 cell lines, representing different MB subtypes. Our results identified a significant number (1,347) of hitherto unrecognized new miRNAs commonly expressed in CM of the 3 cell lines. A significant concordance of ex-miRNA spectra in CM and those expressed intracellularly was observed. Since deregulated miRNA expression is an early event in tumorigenesis, measuring miRNA levels in CSF may also be useful for early detection, which can contribute greatly to the success of treatment.[28] Therefore, in order to use ex-miRNAs as biomarkers for MB, it is important to establish a signature capable of differentiating disease from healthy states. Our pilot microarray screening identified 86 miRNAs exclusively detected in CSF of MB patients but not in control CSF from patients with no brain tumor. We also identified 268 miRNAs that are over-represented and interestingly, only 6 miRNAs under-represented in MB-CSF compared with control CSF. These findings could be of great significance, providing the correlation between expression levels of these miRNAs in CSF of MB patients and their disease states can be established in future studies.
Tumor cell-derived ex-miRNAs are reported to be pro-tumorigenic.[29] Ex-miRNAs can transfer their oncogenic activity to recipient target cells to influence cancer stimulatory activities, thus contributing to the formation of a pre-metastatic niche and promotion of metastasis.[28,30] This exchange of miRNAs between primary tumors and target cells is an interesting and novel dimension to the regulation of a cell phenotype[31-34] and may be particularly important in cancers that have a propensity for dissemination, such as MB. MB includes various subtypes with group 3 and 4 subtypes being clinically distinct with regard to metastasis and prognosis, which may also manifest in a difference in their miRNA spectra. Hence, it was not surprising to find a group of miRNAs that were uniquely over-(60 miRNAs) or under-represented (52 miRNAs) in the CM of the 2 metastasis-related cell lines D283 and D341. More importantly, we identified 4 miRNAs (miR-1290, miR-125a, miR-125b, miR-1298) that were over-represented in MB CSF and significantly enriched in the CM media of the 2 metastasis-related cell lines (D283 and D341). Remarkably, apart from miR-1298, where no functional information is publically available, the 3 other miRNAs (miR-1290, miR-125a, miR-125b) were detected in body fluids of various cancer patients, whereby their increased expression and/or secretion is associated with metastasis of multiple malignancies.[35-39] Consistently, detection of metastasis-related ex-miRNAs in extracellular environment of certain human malignancies, including breast and prostate cancers, were observed in other studies.[40-44] Our observations provide indirect evidence supporting the hypothesis that ex-miRNA are possible facilitators of metastasis by modifying local or distal microenvironments.[45] However, further studies are needed using counter-regulation of key ex-miRNA expression to determine their effect on regulation of motility, migration, and invasion of MB cells.
To the best of our knowledge, this is the first study revealing the spectra of ex-miRNAs in cell CM conditioned by MB cell lines and in CSF of an MB patient. Although the number of samples studied here is very small, our identification of key secreted miRNAs that are specifically enriched in MB-CSF provides a rationale for future investigations. Such investigations, using larger sets of MB samples could lead in the near future to the discovery of CSF-derived miRNA markers, with diagnostic and prognostic significance and ultimately, hopefully also with therapeutic potential.
Acknowledgments
This project was supported by the Swiss Research Foundation Child and Cancer and by “Krebsliga Zürich”.
Financial support and sponsorship
Nil.
Conflicts of interest
There are no conflicts of interest.
REFERENCES
1. Gurney JG, Smith MA, Bunin GR. CNS and miscellaneous intracranial and intraspinal neoplasms In: Ries LA, Smith MA, Gurney JG, Linet M, Tamra T, Young JL, Bunin GR, editors. Cancer Incidence and Survival Among Children and Adolescents: United States SEER Program 1975-1995 Bethesda: National Institutes of Health; 1999. pp. 51-63.
2. Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A, Scheithauer BW, Kleihues P. The 2007 WHO Classification of Tumours of the Central Nervous System. Acta Neuropathol 2007;114:97-109.
3. Tran N, McLean T, Zhang X, Zhao CJ, Thomson JM, O'Brien C, Rose B. MicroRNA expression profiles in head and neck cancer cell lines. Biochem Biophys Res Commun 2007;358:12-7.
4. Calin GA, Croce CM. Chronic lymphocytic leukemia: interplay between noncoding RNAs and protein-coding genes. Blood 2009;114:4761-70.
5. He H, Jazdzewski K, Li W, Liyanarachchi S, Nagy R, Volinia S, Calin GA, Liu CG, Franssila K, Suster S, Kloos RT, Croce CM, de la Chapelle A. The role of microRNA genes in papillary thyroid carcinoma. Proc Natl Acad Sci U S A 2005;102:19075-80.
7. Alsidawi S, Malek E, Driscoll JJ. MicroRNAs in brain metastases: potential role as diagnostics and therapeutics. Int J Mol Sci 2014;15:10508-26.
8. Kosaka N, Iguchi H, Ochiya T. Circulating microRNA in body fluid: a new potential biomarker for cancer diagnosis and prognosis. Cancer Sci 2010;101:2087-92.
9. Chen X, Liang H, Zhang J, Zen K, Zhang CY. Secreted microRNAs: a new form of intercellular communication. Trends Cell Biol 2012;22:125-32.
10. Liang H, Gong F, Zhang S, Zhang CY, Zen K, Chen X. The origin, function, and diagnostic potential of extracellular microRNAs in human body fluids. Wiley Interdiscip Rev RNA 2014;5:285-300.
11. Lawrie CH, Gal S, Dunlop HM, Pushkaran B, Liggins AP, Pulford K, Banham AH, Pezzella F, Boultwood J, Wainscoat JS, Hatton CS, Harris AL. Detection of elevated levels of tumour-associated microRNAs in serum of patients with diffuse large B-cell lymphoma. Br J Haematol 2008;141:672-5.
12. Tanaka M, Oikawa K, Takanashi M, Kudo M, Ohyashiki J, Ohyashiki K, Kuroda M. Down-regulation of miR-92 in human plasma is a novel marker for acute leukemia patients. PLoS One 2009;4:e5532.
13. Huang Z, Huang D, Ni S, Peng Z, Sheng W, Du X. Plasma microRNAs are promising novel biomarkers for early detection of colorectal cancer. Int J Cancer 2010;127:118-26.
14. Heneghan HM, Miller N, Lowery AJ, Sweeney KJ, Newell J, Kerin MJ. Circulating microRNAs as novel minimally invasive biomarkers for breast cancer. Ann Surg 2010;251:499-505.
15. Brase JC, Johannes M, Schlomm T, Fälth M, Haese A, Steuber T, Beissbarth T, Kuner R, Sültmann H. Circulating miRNAs are correlated with tumor progression in prostate cancer. Int J Cancer 2011;128:608-16.
16. Resnick KE, Alder H, Hagan JP, Richardson DL, Croce CM, Cohn DE. The detection of differentially expressed microRNAs from the serum of ovarian cancer patients using a novel real-time PCR platform. Gynecol Oncol 2009;112:55-9.
17. Wang J, Chen J, Chang P, LeBlanc A, Li D, Abbruzzesse JL, Frazier ML, Killary AM, Sen S. MicroRNAs in plasma of pancreatic ductal adenocarcinoma patients as novel blood-based biomarkers of disease. Cancer Prev Res (Phila) 2009;2:807-13.
18. Tsujiura M, Ichikawa D, Komatsu S, Shiozaki A, Takeshita H, Kosuga T, Konishi H, Morimura R, Deguchi K, Fujiwara H, Okamoto K, Otsuji E. Circulating microRNAs in plasma of patients with gastric cancers. Br J Cancer 2010;102:1174-9.
19. Hu Z, Chen X, Zhao Y, Tian T, Jin G, Shu Y, Chen Y, Xu L, Zen K, Zhang C, Shen H. Serum microRNA signatures identified in a genome-wide serum microRNA expression profiling predict survival of non-small-cell lung cancer. J Clin Oncol 2010;28:1721-6.
20. Baraniskin A, Kuhnhenn J, Schlegel U, Maghnouj A, Zöllner H, Schmiegel W, Hahn S, Schroers R. Identification of microRNAs in the cerebrospinal fluid as biomarker for the diagnosis of glioma. Neuro Oncol 2012;14:29-33.
21. Akers JC, Ramakrishnan V, Kim R, Skog J, Nakano I, Pingle S, Kalinina J, Hua W, Kesari S, Mao Y, Breakefield XO, Hochberg FH, Van Meir EG, Carter BS, Chen CC. MiR-21 in the extracellular vesicles (EVs) of cerebrospinal fluid (CSF): a platform for glioblastoma biomarker development. PLoS One 2013;8:e78115.
22. Teplyuk NM, Mollenhauer B, Gabriely G, Giese A, Kim E, Smolsky M, Kim RY, Saria MG, Pastorino S, Kesari S, Krichevsky AM. MicroRNAs in cerebrospinal fluid identify glioblastoma and metastatic brain cancers and reflect disease activity. Neuro Oncol 2012;14:689-700.
23. von Bueren AO, Shalaby T, Rajtarova J, Stearns D, Eberhart CG, Helson L, Arcaro A, Grotzer MA. Anti-proliferative activity of the quassinoid NBT-272 in childhood medulloblastoma cells. BMC Cancer 2007;7:19.
24. Kosaka N, Iguchi H, Yoshioka Y, Takeshita F, Matsuki Y, Ochiya T. Secretory mechanisms and intercellular transfer of microRNAs in living cells. J Biol Chem 2010;285:17442-52.
25. Lukiw WJ, Alexandrov PN, Zhao Y, Hill JM, Bhattacharjee S. Spreading of Alzheimer's disease inflammatory signaling through soluble micro-RNA. Neuroreport 2012;23:621-6.
26. Turchinovich A, Weiz L, Langheinz A, Burwinkel B. Characterization of extracellular circulating microRNA. Nucleic Acids Res 2011;39:7223-33.
27. Snuderl M, Batista A, Kirkpatrick ND, Ruiz de Almodovar C, Riedemann L, Walsh EC, Anolik R, Huang Y, Martin JD, Kamoun W, Knevels E, Schmidt T, Farrar CT, Vakoc BJ, Mohan N, Chung E, Roberge S, Peterson T, Bais C, Zhelyazkova BH, Yip S, Hasselblatt M, Rossig C, Niemeyer E, Ferrara N, Klagsbrun M, Duda DG, Fukumura D, Xu L, Carmeliet P, Jain RK. Targeting placental growth factor/neuropilin 1 pathway inhibits growth and spread of medulloblastoma. Cell 2013;152:1065-76.
28. Cortez MA, Bueso-Ramos C, Ferdin J, Lopez-Berestein G, Sood AK, Calin GA. MicroRNAs in body fluids - the mix of hormones and biomarkers. Nat Rev Clin Oncol 2011;8:467-77.
29. Hannafon BN, Ding WQ. Intercellular Communication by Exosome-Derived microRNAs in Cancer. Int J Mol Sci 2013;14:14240-69.
30. Wang J, Zhang KY, Liu SM, Sen S. Tumor-associated circulating microRNAs as biomarkers of cancer. Molecules 2014;19:1912-38.
31. Mittelbrunn M, Sánchez-Madrid F. Intercellular communication: diverse structures for exchange of genetic information. Nat Rev Mol Cell Biol 2012;13:328-35.
32. Frampton AE, Krell J, Kazemier G, Giovannetti E. Serum miR-1290 as a marker of pancreatic cancer - letter. Clin Cancer Res 2013;19:5250-1.
33. Hammond J, Johnson HM, Varas R, Ward CG. A qualitative comparison of paper flowsheets vs a computer-based clinical information system. Chest 1991;99:155-7.
34. Huang X, Yuan T, Liang M, Du M, Xia S, Dittmar R, Wang D, See W, Costello BA, Quevedo F, Tan W, Nandy D, Bevan GH, Longenbach S, Sun Z, Lu Y, Wang T, Thibodeau SN, Boardman L, Kohli M, Wang L. Exosomal miR-1290 and miR-375 as prognostic markers in castration-resistant prostate cancer. Eur Urol 2015;67:33-41.
35. Korzeniewski N, Tosev G, Pahernik S, Hadaschik B, Hohenfellner M, Duensing S. Identification of cell-free microRNAs in the urine of patients with prostate cancer. Urol Oncol 2015;33:16.e17-22.
36. Le MT, Teh C, Shyh-Chang N, Xie H, Zhou B, Korzh V, Lodish HF, Lim B. MicroRNA-125b is a novel negative regulator of p53. Genes Dev 2009;23:862-76.
37. Sand M, Skrygan M, Sand D, Georgas D, Hahn SA, Gambichler T, Altmeyer P, Bechara FG. Expression of microRNAs in basal cell carcinoma. Br J Dermatol 2012;167:847-55.
38. Sun X, Song Y, Tai X, Liu B, Ji W. MicroRNA expression and its detection in human supraglottic laryngeal squamous cell carcinoma. Biomed Rep 2013;1:743-746.
39. Sun YM, Lin KY, Chen YQ. Diverse functions of miR-125 family in different cell contexts. J Hematol Oncol 2013;6:6.
40. Tang F, Zhang R, He Y, Zou M, Guo L, Xi T. MicroRNA-125b induces metastasis by targeting STARD13 in MCF-7 and MDA-MB-231 breast cancer cells. PLoS One 2012;7:e35435.
41. Wang H, Tan G, Dong L, Cheng L, Li K, Wang Z, Luo H. Circulating MiR-125b as a marker predicting chemoresistance in breast cancer. PLoS One 2012;7:e34210.
42. Cheng HH, Mitchell PS, Kroh EM, Dowell AE, Chéry L, Siddiqui J, Nelson PS, Vessella RL, Knudsen BS, Chinnaiyan AM, Pienta KJ, Morrissey C, Tewari M. Circulating microRNA profiling identifies a subset of metastatic prostate cancer patients with evidence of cancer-associated hypoxia. PLoS One 2013;8:e69239.
43. Kosaka N, Iguchi H, Hagiwara K, Yoshioka Y, Takeshita F, Ochiya T. Neutral sphingomyelinase 2 (nSMase2)-dependent exosomal transfer of angiogenic microRNAs regulate cancer cell metastasis. J Biol Chem 2013;288:10849-59.
44. Zhang Y, Yang P, Wang XF. Microenvironmental regulation of cancer metastasis by miRNAs. Trends Cell Biol 2014;24:153-60.
45. Salido-Guadarrama I, Romero-Cordoba S, Peralta-Zaragoza O, Hidalgo-Miranda A, Rodríguez-Dorantes M. MicroRNAs transported by exosomes in body fluids as mediators of intercellular communication in cancer. Onco Targets Ther 2014;7:1327-38.
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Shalaby T, Fiaschetti G, Baulande S, Gerber NU, Baumgartner M, Grotzer MA. Detection and quantification of extracellular microRNAs in medulloblastoma. J Cancer Metastasis Treat 2015;1:67-75. http://dx.doi.org/10.4103/2394-4722.157068
AMA Style
Shalaby T, Fiaschetti G, Baulande S, Gerber NU, Baumgartner M, Grotzer MA. Detection and quantification of extracellular microRNAs in medulloblastoma. Journal of Cancer Metastasis and Treatment. 2015; 1: 67-75. http://dx.doi.org/10.4103/2394-4722.157068
Chicago/Turabian Style
Shalaby, Tarek, Giulio Fiaschetti, Sylvain Baulande, Nicolas U. Gerber, Martin Baumgartner, Michael A. Grotzer. 2015. "Detection and quantification of extracellular microRNAs in medulloblastoma" Journal of Cancer Metastasis and Treatment. 1: 67-75. http://dx.doi.org/10.4103/2394-4722.157068
ACS Style
Shalaby, T.; Fiaschetti G.; Baulande S.; Gerber NU.; Baumgartner M.; Grotzer MA. Detection and quantification of extracellular microRNAs in medulloblastoma. J. Cancer. Metastasis. Treat. 2015, 1, 67-75. http://dx.doi.org/10.4103/2394-4722.157068
About This Article
Copyright
Data & Comments
Data
 Cite This Article 2 clicks
Cite This Article 2 clicks











Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.