Profiling of circulating tumor cells in liquid biopsies from metastatic cancer patients
Abstract
Aim: Circulating tumor cells (CTCs) are crucial to tumor metastasis and valuable for prediction of clinical outcome in patients with solid tumors. Here, the authors aimed to establish a method for enumeration and characterization of CTCs from liquid biopsies. Methods: Peripheral blood mononuclear cells (PBMCs) were separated from blood samples from patients with metastatic cancer using Ficoll-Hypaque gradients and cultured to isolate and enumerate CTCs. Cultured CTCs were morphologically characterized by light and phase contrast microscopy. The tumorigenicity of Ficoll-Hypaque-separated PBMCs was examined, in addition to their expression of mRNA metastasis markers. Results: CTCs were isolated in culture and enumerated by counting under phase contrast microscopy, demonstrating that 0.01-0.04% of total PBMCs were CTCs. CTCs were dormant, with large, oval-shaped, spiky morphology. PBMCs obtained from liquid biopsies exhibited anchorage-independent growth, forming numerous colonies in soft agar assays. Molecular profiling demonstrated expression of several metastatic genes, but not of cadherin 1 (encoding the adhesion protein), in all patients. Conclusion: The authors successfully isolated, enumerated, and characterized CTCs from liquid biopsies of metastatic cancer patients. This study has potential to facilitate the development of new diagnostic and therapeutic methods using liquid biopsies, for application in metastatic cancers.
Keywords
Introduction
Cancer cells are invasive and can metastasize; once metastasis has occurred, disease is usually incurable. Recent technological developments have enabled evaluation of the metastatic potential of cells using “liquid biopsies”,[1,2] noninvasive tests for circulating tumor cells (CTCs), which are present even at early stages of disease.[3] Additionally, molecular profiling and enumeration of CTCs can predict their metastatic potential, thereby facilitating improved treatment and prognosis.[3] Such testing has potential to revolutionize personalized cancer medicine by avoiding biopsies, since it captures cancer cells that tumors shed into the circulation, rather than sampling the tumor itself. Liquid biopsies permit repeated, noninvasive sample collection from cancer patients that can be profiled for gene expression, allowing clinicians to choose specific therapies targeting particular mutations. Ultimately, liquid biopsies could become a routine cancer screening method, alongside current methods, such as mammograms and colonoscopies.[4,5] Liquid biopsies could be useful in the majority of invasive solid tumors, including breast, colon, lung, prostate, and pancreatic cancers.
Isolation and enumeration of CTCs is difficult because of their low numbers, size, and heterogeneity. Several methods have been established for isolation and enumeration of CTCs from liquid biopsies, including the Ficoll-Hypaque gradient method,[6] flow cytometry,[7,8] the CELLSEARCH® system,[9] isolation by size of epithelial tumor cells (ISET),[10,11] and microfluidic systems.[12] The CELLSEARCH® system is primarily based on selection of EpCAM- and cytokeratin-positive CTCs, and has been approved by the Food and Drug Administration (FDA) and used in several clinical studies;[13,14] however, one study reported that CTCs which have undergone epithelial mesenchymal transition (EMT) exhibit down-regulation of epithelial markers, including EpCAM and keratin,[15] and such CTCs would be missed using this technology. Similarly, in some cases, CTCs from HER2-positive metastatic breast cancer patients include EpCAM-negative cells.[15] Hence, EpCAM-independent methods may enable improved detection of CTCs in certain cancer patients. Advances in microfluidic technologies, biomaterials, and molecular profiling have led to rapid growth and interest in achieving liquid biopsies for cancer diagnosis and treatment. CTCs are primarily characterized and identified by their morphology and immunostaining pattern with specific antibodies; however, the heterogeneity of CTC’s is a major obstacle to their isolation, identification, and characterization from liquid biopsies.
Chemotherapy is a standard mode of treatment for all cancers. CTC levels are determined before and after rounds of chemotherapy. In the majority of cases of non-metastatic breast cancer, reduction in the number of CTCs is observed after the first round of chemotherapy; however, CTCs can be resistant to therapy, leading to administration of increasing doses in subsequent rounds of chemotherapy. In one study, patients with > 80 CTCs in 7.5 mL of blood died within one year of diagnosis due to metastases.[16] In another study of patients with metastatic breast cancer, when CTCs continued to be detected after the first round of chemotherapy, there was rapid progression of the disease; it was suggested that, in such cases, it would be preferable to opt for alternative treatment, rather than continuing with the same chemotherapy.[17] A review of CTCs in patients with breast cancer in 2013 considered the clinical application of CTCs in breast cancer therapy and assessed the use of HER2 treatment in HER2-negative breast cancer patients, selected on the basis of CTC detection.[13,14] Rack et al.[18] concluded that there is an independent prognostic relevance of CTC determination both before and after adjuvant chemotherapy in a study of > 2,000 patients with primary breast cancer. Bidard et al.[13] also studied the clinical utility of CTCs in metastatic breast cancer, mainly focusing on first and second line treatments.
Completion of the human genome project and the advent of molecular profiling has led to an understanding of the genetic profiles of cancer cells, including CTCs.[19]MMP1, MMP2, MMP9, VEGFA, MT-CO2, ICAM1, CD44, and PROM1 are major genes involved in the processes of invasion and metastasis. Moreover, cadherin-1, N-cadherin, fibronectin, and integrin β are adhesion molecules involved in the metastatic process. During invasion and metastasis, CTCs frequently undergo the process of EMT, which involves several genes including TWIST1 and TWIST2,[20] in combination with TGFB1, WNT, and NOTCH2. The majority of CTCs isolated from breast cancer patients express EMT markers, including ETV5, NOTCH1, SNAIL, TGFB1, ZEB1, and ZEB2.[21] Keratin analysis is also a major molecular test in cancer, while CK19 and TP53 mutations are frequently found in CTCs from triple-negative breast cancer patients,[22] and are a factor in the progression of the disease to the triple-negative stage. Patients with breast cancer expressing CK19, SCGB2A2, and ERBB2 showed poor survival rates.[22] Riethdrof et al.[23] demonstrated that HER2+ CTCs can be identified in HER2- breast cancer patients, leading to consideration of revision of ongoing treatment, with trastuzumab-based therapy applied to patients with HER2+ CTCs and HER2- primary tumors, demonstrating that CTCs can assist in determining the changing course of disease in a timely manner and have potential to determine the metastatic state of breast cancer.
CTCs are present in very low numbers in whole blood and are difficult to identify and characterize. The primary aims of the present study were to establish a simple protocol for the isolation, identification, and enumeration of CTCs from various metastatic cancer patients and to molecularly profile genes involved in the processes of invasion and metastasis, and to evaluate the potential clinical utility of liquid biopsies in the treatment of advanced stage cancer.
Methods
Materials
Low-glucose Dulbecco’s modified Eagle’s Medium (DMEM), penicillin/streptomycin (PenStrep), phosphate-buffered saline (PBS), trypsin EDTA, erythrosin B, and colchicine, were purchased from HiMedia (Mumbai, India); fetal bovine serum (FBS) from GIBCO BRL (Carboside, MA); Trizol reagent, cDNA preparation kits, and agarose from Invitrogen (Carlsbad, CA, USA); and histopaque and primers for KRT18, KRT19, PROM1, CD44, CXCR4, NOTCH2, VEGFA, MMP1, MMP2, MMP9, ICAM1, CDH1, KCNH2, and ACTB from Sigma Chemicals, USA.
Sample collection
A total of eight metastatic cancer patients recruited by oncology clinic of Jaslok Hospital and Research Centre, Mumbai India (three breasts, two ovarian, two prostate, and one nasopharyngeal cancer) and five healthy individuals were included in this study. All patients had stage IV disease, with invasive and metastatic cancer. All tumors were histopathologically proven to have metastatic potential. Fresh blood samples (10 mL) were collected from each metastatic cancer patient and healthy control individual in sterile EDTA vacutainers with proper consent from the patients, according to the ethical committee guidelines of Jaslok Hospital and Research Center, Mumbai, India, and sent to the tissue culture laboratory of the Molecular Medicine and Biology Department.
Culture of peripheral blood mononuclear cells for isolation of CTCs
Plasma rich in peripheral blood mononuclear cells (PBMCs) was separated from blood samples from cancer patients and healthy individuals by allowing blood to stand for approximately 1 h. We obtained approximately 2 mL of PBMC-rich plasma from 10 mL of blood. Next, PBMCs were isolated by Ficoll-Hypaque gradient centrifugation as follows. Briefly, 300 µL aliquots of PBMC-rich plasma were mixed with 300 µL of 1 × PBS and this mixture was layered onto 300 µL of Ficoll-Hypaque solution. We prepared two tubes each for cancer patients and healthy individuals for our culture study. Tubes were centrifuged at 3,000 g for 30 min to obtain a middle layer containing PBMCs. The PBMCs isolated from the two tubes were pooled together, washed with 1 × PBS, and suspended in 4 mL of RPMI growth media, supplemented with 10% FBS, 1% PenStrep, L-glutamine, and vitamin C. Tissue culture dishes (65 mm) containing PBMCs were incubated in a 5% CO2 incubator at 37 °C. After 24 h, non-adherent cells were removed and the remaining adherent cells fed with 4 mL of DMEM growth medium supplemented with 10% FBS + PenStrep + L-glutamine, which were changed twice a week. Cultures were observed and photographed every day for 30 days under a phase contrast microscope (model AXiovert 40CFL from Carl Zeiss), equipped with TS view software (Tucsen Imaging, Fuzhou, and PR China) and images were captured and analyzed to determine and record the morphology of adherent cells. Adherent cells that appeared CTC-like by phase contrast microscopy were counted manually under a phase contrast microscope to calculate the number of cells present in each metastatic cancer patient and were then fixed in 50% methanol, stained with Giemsa, and examined by light microscopy (AXiovert 40CFL, Carl Zeiss) to determine their general morphological features.
Anchorage-independent soft agar assay
PBMCs were isolated from metastatic cancer patients and healthy individuals and their tumorigenic potential determined using soft agar assays. A total of 3 × 103 PBMCs per individual were layered on 0.4% soft agar in DMEM growth medium in 65 mm dishes. The plates were then incubated at 37 °C with 5% CO2 for 2 weeks. Emerging colonies were observed under a phase contrast microscope and photographed.
Molecular markers in PBMCs (liquid biopsies) and cultured CTCs
Total RNA was extracted from all patient’s PBMCs and isolated cultured CTCs from Ovarian, prostate and CNS cancer patients using the Trizol method. RNA was then reverse transcribed to cDNA using the Applied Biosystems High Capacity cDNA Kit (Applied Biosystems, USA). PCR was carried out using specific forward and reverse primers with defined annealing temperatures [Table 1]. For all genes, PCR reactions were performed at 95 °C for 5 min; followed by 40 cycles at 95 °C for 30 s, at the respective annealing temperatures [Table 1] for 30 s, and at 72 °C for 30 s; with a final extension at 72 °C for 7 min. Amplicon sizes were checked by 2% agarose gel electrophoresis, followed by using Ethidium Bromide dye and visualization using a gel documentation system (Alpha Imager HP from Cell Bioscience) and photography.
Forward and reverse primer sequences used for respective molecular markers with their annealing temperature and size
| Serial No. | Name | Primer sequence (5’-3’) | Annealing (°C) | Size (bp) |
|---|---|---|---|---|
| 1 | ACTIN F | GACTACCTCATGAAGATC | 55 | 512 |
| 2 | ACTIN R | GATCCACATCTGCTGCAA | 55 | 512 |
| 3 | KERATIN F | GAGATCGAGGCTCTCAAGGA | 55 | 357 |
| 4 | KERATIN R | CAAGCTGGCCTTCAGATTTC | 55 | 357 |
| 5 | CD44 F | CAACCCTACTGATGATGACG | 60 | 302 |
| 6 | CD44 R | GGATGCCAAGATGATCAGCC | 60 | 302 |
| 7 | CXCR4 F | GGACCTGTGGCCAAGTTCTTAGTT | 60 | 273 |
| 8 | CXCR4 R | ACTGTAGGTGCTGAAATCAACCCA | 60 | 273 |
| 9 | NOTCH-2 F | ACTTCCTGCCAAGCATTCC | 60 | 278 |
| 10 | NOTCH-2 R | GTCCATGTCTTCAGTGAGAAC | 60 | 278 |
| 11 | CD133 F | ACCTGCGTAATCCCATCT | 60 | 340 |
| 12 | CD133 R | TTGTCCGACCAGTTCTTC | 60 | 340 |
| 15 | VEGFR F | GAAGTGGTGAAGTTCATGGATGTC | 62 | 422 |
| 16 | VEGFR R | CGATCGTTCTGTATCAGTCTTTCC | 62 | 422 |
| 17 | MMP1 F | CTGAAGGTGATGAAGCAGCC | 55 | 427 |
| 18 | MMP1 R | AGTCCAAGAGAATGGCCGAG | 55 | 427 |
| 19 | MMP2 F | GCGACAAGAAGTATGGCTTC | 58 | 390 |
| 20 | MMP2 R | TGCCAAGGTCAATGTCAGGA | 58 | 390 |
| 21 | MMP9 F | CGCAGACATCGTCATCCAGT | 64 | 405 |
| 22 | MMP9 R | GGATTGGCCTTGGAAGATGA | 64 | 405 |
| 23 | E -Cadherin F | TGCTCTTGCTGTTTCTTCGG | 60 | 422 |
| 24 | E -Cadherin R | TGCCCCATTCGTTCAAGTAG | 60 | 422 |
| 25 | I -CAM1 F | AGGCCACCCCAGAGGACAAC | 58 | 405 |
| 26 | I -CAM1 R | CCCATTATGACTGCGGCTGCTA | 58 | 405 |
Results
Isolation of CTCs from PBMCs from metastatic patients
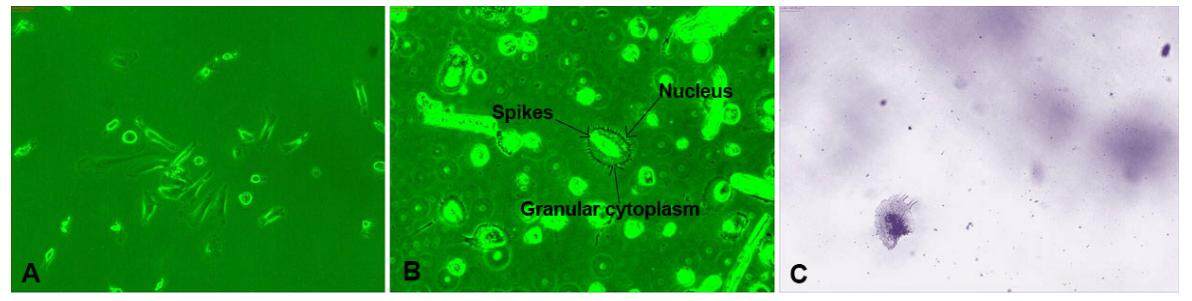
PBMCs were isolated from metastatic cancer patients, counted using a hemocytometer, and 3 × 105 cells were cultured for 24 h, followed by culture only of adherent cells. After 15-20 days of incubation, we observed circular cells with spikes on their circumference [Figure 1], which were metastatic tumor cells that had been circulating in the blood of the cancer patients. We considered these cells to be CTCs and examined their morphology and other characteristics daily by phase contrast microscopy. CTCs were large spherical cells, with spikes, single nuclei, and granular cytoplasm, which differed from other cell types [Figure 1], clearly indicating that the cells observed in samples from metastatic patient were CTCs involved in cancer metastasis. CTCs isolated from all metastatic cancer patients were morphologically similar, with no differences in samples from patients with different types of cancer [Figure 1]. No such cells were observed in PBMCs from healthy individuals cultured simultaneously.
Figure 1. Morphology of circulating tumor cells (×20). A and B are phase contrast microscopy and C is Giemsa stained
To determine the general morphological feature of CTCs, we stained them with Giemsa, revealing that these large cells had distinct nuclei, with granulated cytoplasm at the periphery of the nucleus [Figure 1]. CTCs were dormant, and did not multiply for several months.
Enumeration of CTCs in metastatic cancer patients
Enumeration of CTCs in metastatic cancer patients was a major aim of this study. After 30 days of culture of adherent cells emerging from PBMCs, CTCs were clearly visible in culture dishes [Figure 1] and were counted manually under phase contrast microscopy. In samples from the three breast cancer patients, CTC counts ranged from 120 to 160 cells (average = 145 cells; percentage of total cells plated = 0.045%); from the two patients with prostate cancer, the count ranged from 120 to 160 (average 140 cells; percentage of total cells plated, 0.042%); samples from the two ovarian cancer patients yielded 90-120 CTCs (average = 105; percentage of total cells plated = 0.032%); and from the single nasopharyngeal cancer sample we obtained 50 CTCs (percentage of total cells = 0.015%). All experiments were performed in duplicate. This cell counting process was very consistent and a successful straightforward method to enumerate CTCs.
Anchorage-independent soft agar assays
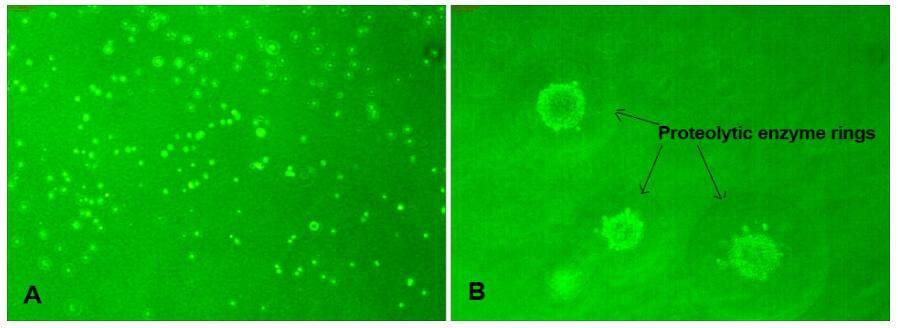
Anchorage-independent soft agar assays were performed to determine the tumorigenic potential of PBMCs isolated from metastatic cancer patients and healthy individuals. No colonies were observed in samples from healthy individuals [Figure 2A]. In contrast, several large colonies grew in agar plates containing PBMCs isolated from all metastatic cancer patients [Figure 2B]. Hence PBMC samples from metastatic cancer patients clearly contained a tumorigenic population, whereas those from healthy individuals did not. We also observed the formation of circular rings around each colony, indicating that the cells secreted proteolytic enzymes [Figure 2B]. This assay only indicates the basic cancer phenotype of cells; tumorigenic potential can be confirmed by “in vivo transformation assays” using nude mice, or by PCR determining their molecular phenotypes. We confirmed the tumorigenicity of the cells by molecular analysis.
Molecular markers in isolated cultured CTCs
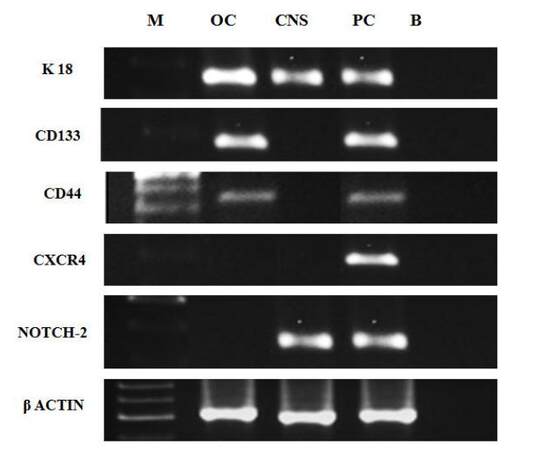
To determine the molecular cancer phenotypes of isolated CTCs, we harvested cells after 30 days of culture of PBMCs from metastatic cancer patients and determined the expression of specific mRNAs involved in metastasis in these cells by RT-PCR. We studied the expression of only five specific genes: KRT18, PROM1, CD44, CXCR4, and NOTCH2, in addition to the ACTB housekeeping gene in these samples, due to the limited quantities of mRNA available. We obtained CTCs from ovarian, prostate, and central nervous system (CNS) cancers for this analysis. We found that ovarian cancer CTCs expressed KRT18, PROM1, and CD44; however, CXCR4 and NOTCH2 were not expressed in CTCs obtained from this cancer type [Figure 3]. CTCs from CNS cancer exhibited expression of KRT18 and NOTCH2 and absence of expression of PROM1, CD44, and CXCR4, whereas prostate cancer CTCs expressed all five genes [Figure 3]. Overall our results indicated that the molecular profiles of CTCs varied according to the type of cancer; therefore, we suggest that individual profiling of metastatic cancer patients will be essential for management of therapy in these patients.
Molecular markers in PBMCs from metastatic cancer patients (liquid biopsies)
We also examined molecular markers present in whole plasma PBMCs (liquid biopsies), from patients with metastatic breast (n = 3), ovarian (n = 1), nasopharyngeal (n = 1), and prostate (n = 1) cancer. We analyzed expression of 11 genes related to metastasis: KRT18, CD44, PROM1, CXCR4, NOTCH2, MMP1, MMP2, MMP9, KCNH2, ICAM1 and CADH1. KRT18, CD44, PROM1, CXCR4, and NOTCH2 were expressed by PBMCs from all patients [Figure 4] indicating that they were present in all cancer patients and have roles in the metastatic process.
Figure 4. Molecular profiling of liquid biopsies (PBMC) of metastatic cancer patients. PBMC: peripheral blood mononuclear cell
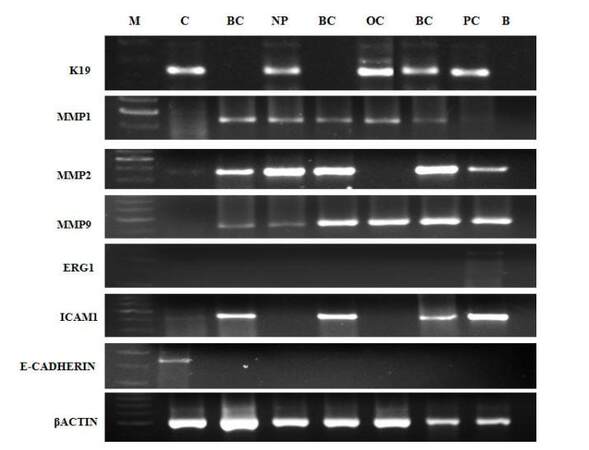
The expression of KRT19, MMP1, MMP2, MMP9, ICAM, and CDH1 was studied in samples from three breast cancer patients, one ovarian cancer patient, one nasopharyngeal cancer patient, and one prostate cancer patient, and the results are shown in Figure 5. KRT19 was expressed in the ovarian, nasopharyngeal, and prostate cancer samples, whereas its expression was completely absent in samples from all three breast cancer patients; and the metastatic genes, MMP1, MMP2, and MMP9, were expressed in all patients studied, confirming the metastatic potential of these cells. The MMP2 gene was highly expressed in breast and nasopharyngeal cancer, whereas KCNH2, which is specific for prostate cancer, was faintly expressed only in the sample from the prostate cancer patient and its expression was absent from all other samples. The ICAM gene, which encodes a molecule involved in cellular adhesion, was expressed in breast and prostate cancer samples, but expression was absent from ovarian and nasopharyngeal cancer samples; in contrast, the expression of another gene, CADH1, encoding the adhesion molecule (cadherin 1), was completely absent from the samples from all four types of cancer patients, indicating that cells in liquid biopsies from metastatic cancer patients were in the EMT phase, in which they lose their adhesive properties and become free to move in the blood stream, which may also explain the high metastatic potential of these cells.
Discussion
In their early stages, the majority of cancers are asymptomatic, whereas they exhibit rapid growth rates later in disease progression; therefore, it is difficult to treat advanced-stage cancer patients, due to spread of the disease to various organs and tissues.[24] CTCs are shed by primary tumors into the vasculature and circulate in the blood of cancer patients;[25] however, they are not easily identified, due to their very limited numbers. Pantel et al.[26] reviewed the importance of CTCs, including the biological properties of metastatic cells involved in cancer progression, and stated that CTC-derived cell lines could be used to develop new therapeutic targets, and for drug screening. Several investigators have described the development of in vivo and in vitro methods for the isolation of CTCs in larger numbers from metastatic cancer patients, for use in research into the role of CTCs in cancer progression,[27-29] and with the aim of targeting such cells to cure cancer; hence CTCs are a topic of intense discussion among oncologists.[25]
Many techniques have been developed and are under continuous improvement to enhance the efficacy of CTC isolation and enumeration.[16] Comparison of the OncoQuick density gradient method with Ficoll-Hypaque density gradients for separation of tumor cells from bone marrow and peripheral blood aspirates indicated that CTCs can be easily aspirated by both methods and analyzed further to determine their presence and quantity.[30] In the present study, we used the simple and cost-effective Ficoll-Hypaque density gradient method to separate PBMCs from whole plasma of cancer patients. These cells were then cultured to isolate CTCs, which were identified by their morphology (i.e. large oval shape, with large nuclei and peripheral spikes) [Figure 1]. Similar morphology was first reported by Gascoyne et al.[31] It is likely that the spike-like processes allow CTCs to adhere to the surface of basement membranes during the process of invasion. We confirmed the phenotype of these cells by studying cancer-associated genes in isolated CTCs and from liquid biopsies [Figures 3 and 4].
The presence of CTCs in an advanced metastatic cancer patient was first reported in 1869 by Ashworth;[32] however, limited information is available regarding the numbers of CTCs in the blood of patients with different stages and types of cancer,[33] making proficient isolation of CTCs in a viable and intact state challenging. The CELLSEARCH® system[34] is the only method approved by the US FDA for clinical use in metastatic breast, colorectal, and prostate cancer.[13,17,35] Patients with > 80 CTCs in 7.5 mL of blood were reported to die within one year of diagnosis from metastatic disease,[16] and CTCs continue to be detected after the first round of chemotherapy in some breast cancer patients with rapid progression of the disease to metastasis;[17] it has been suggested that, in such cases, it may be preferable to opt for an alternative treatment with novel therapeutic agents, rather than continuing with the same chemotherapy.
In the present study, we established a simple method of isolation and enumeration of CTCs from metastatic cancer patients. PBMCs were cultured in 65-mm dishes and numbers of CTC-like cells were counted manually under a phase contrast microscope. We observed that there were more CTCs in patients with final-stage, compared with early-stage, metastatic cancer. We found approximately 50-160 CTCs in the majority of cancer patients, representing 0.01-0.04% of overall PBMCs. All three breast cancer patients and one prostate cancer patient were determined to be in the late stages of metastatic disease, according to CTC enumeration, which was confirmed by molecular profiling. Hence, our results indicate that enumeration and molecular profiling of CTCs can distinguish early- and later-stage disease in metastatic cancer patients. Moreover, the values obtained by enumeration of CTCs correlated well with those from previous reports.[13,14]
The CELLSEARCH® system, is primarily selective for EpCAM- and cytokeratin-positive CTCs;[34] however, Lianidou et al.[15] demonstrated that CTCs which have undergone EMT exhibit down-regulation of epithelial markers, including EpCAM and K19 and hence such cells will be missed by this technology. Similarly, some cases of HER2-positive metastatic breast cancer have EpCAM-negative CTCs. Hence, additional markers should be considered in the development of methods to isolate and enumerate CTCs in metastatic cancer. Our study also demonstrated that KRT19 and CADH1 were not expressed in liquid biopsies from any breast cancer patients [Figure 5]. Moreover, the observed absence of CADH1 expression indicated that CTCs obtained from all cancer patients were in EMT. As the method we used was based on counting intact cultured CTCs by phase contrast microscopy, the possibility of missing CTCs was small.
The soft agar colony formation assay is a common method to monitor anchorage-independent growth in semi-solid agar during 3-4 week incubation by manual counting of colonies under phase contrast microscopy. This method is considered one of the most accurate and sensitive in vitro assays for detection of malignant transformation of cells and confirmation of the phenotype of tumor cells with metastatic potential.[36] In this study, we found that all PBMCs isolated from metastatic cancer patients resulted in significant numbers of colonies on soft agar, indicating their high tumorigenic potential. There were rings around each colony [Figure 2], indicating the activity of secretory proteases from these cells, which are primarily responsible for the degradation of basement membranes during invasion and metastasis.[37]
Molecular profiling of tumor cells has become important for understanding genes involved in cancer development,[38] and provides insights into the selection of tumor cells and resistance mechanisms in patients undergoing systemic therapies; therefore, molecular profiling of liquid biopsies or isolated CTCs, to determine the major genes involved in the processes of invasion and metastasis, was a primary aim of this study. We performed molecular profiling of isolated CTCs as well as liquid biopsies obtained from metastatic cancer patients. The results indicated that CTCs and liquid biopsies from all cancer patients expressed KRT18, PROM1, CD44, CXCR4, and NOTCH2 genes [Figures 3 and 4], demonstrating that the malignant phenotypes of CTCs and liquid biopsies can be evaluated by molecular profiling. We also observed that KRT19 was not expressed down-regulated in all breast cancer patients, possibly due to EMT of these cells during the invasive process, and suggesting a need for aggressive treatment for these patients. MMP1, MMP2, and MMP9 are metastatic genes involved in the invasion of tumor cells into underlying epithelium.[39] This is the first report that liquid biopsies from all cancer patients tested express high levels of MMP1, MMP2, and MMP9, providing a potential new avenue for treatment of metastatic cancers by suppression of these genes; however, these results require confirmation and further investigation. PBMC from five normal individual were tested and found to be negative for expression of cancer-related genes.
Adhesion molecules play important roles in the migration of tumor cells to distant organs.[40] Cadherin 1 is primarily responsible for adherence of cells to surfaces and reduced mobility of tumor cells; however, due to down-regulation of cadherin 1, metastatic cells are highly mobile and can easily travel to distant organs in short periods of time.[41] Our results indicated that the gene encoding cadherin 1 (CADH1) was undetected in all cancers studied, indicating that all CTCs obtained from metastatic cancer patients were in EMT.
Overall, this is the first report of molecular profiling of several cancer genes in isolated CTCs and liquid biopsies from metastatic cancer patients. We suggest that liquid biopsies can be profiled for the metastatic genes identified in this study as potentially useful for the monitoring and treatment of metastatic cancer patients, with the aim of achieving improved outcomes. We realize that additional tests, including immunofluorescence, will be required to fully characterize the cells we have observed in metastatic patients’ blood. We plan to perform these studies in the near future.
In conclusion, the present study describes a simple protocol for the isolation and enumeration of CTCs from liquid biopsies and the morphological and molecular characterization of CTCs from metastatic cancer patients. Molecular profiling of CTCs and whole liquid biopsies clearly indicated the expression of metastatic genes and adhesion molecules involved in the metastatic process; thus molecular profiling for these genes could enable the establishment of improved diagnosis and treatment protocols for last stage cancer using liquid biopsies.
Financial support and sponsorship
Nil.
Conflicts of interest
There are no conflicts of interest.
Patient consent
Patient consent was obtained from the patients.
Ethics approval
The study has been approved by ethical committee guidelines of Jaslok Hospital and Research Center.
REFERENCES
1. Mocellin S, Keilholz U, Rossi CR, Nitti D. Circulating tumor cells: the 'leukemic phase' of solid cancers. Trends Mol Med 2006;12:130-9.
2. Marrinucci D, Bethel K, Kolatkar A, Luttgen MS, Malchiodi M, Baehring F, Voigt K, Lazar D, Nieva J, Bazhenova L, Ko AH, Korn WM, Schram E, Coward M, Yang X, Metzner T, Lamy R, Honnatti M, Yoshioka C, Kunken J, Petrova Y, Sok D, Nelson D, Kuhn P. Fluid biopsy in patients with metastatic prostate, pancreatic and breast cancers. Phys Biol 2012;9:016003.
3. Pantel K, Brakenhoff RH, Brandt B. Detection, clinical relevance and specific biological properties of disseminating tumour cells. Nat Rev Cancer 2008;8:329-40.
4. Witzig TE, Bossy B, Kimlinger T, Roche PC, Ingle JN, Grant C, Donohue J, Suman VJ, Harrington D, Torre-Bueno J, Bauer KD. Detection of circulating cytokeratin-positive cells in the blood of breast cancer patients using immunomagnetic enrichment and digital microscopy. Clin Cancer Res 2002;8:1085-91.
5. Takeuchi H, Kitagawa Y. Circulating tumor cells in gastrointestinal cancer. J Hepatobiliary Pancreat Sci 2010;17:577-82.
6. Racila E, Euhus D, Weiss AJ, Rao C, McConnell J, Terstappen LW, Uhr JW. Detection and characterization of carcinoma cells in the blood. Proc Natl Acad Sci USA 1998;95:4589-94.
7. Takao M, Takeda K. Enumeration, characterization, and collection of intact circulating tumor cells by cross contamination-free flow cytometry. Cytometry A 2011;79:107-17.
8. Hong B, Mace M, Crowder R, Coleman T, He W, Szczepanski F, Feczko J, Lyzak J. Metastatic breast cancer detection and therapy monitoring using folate-targeting flow cytometry. J Clin Oncol 2011;29:23.
9. Allard WJ, Matera J, Miller MC, Repollet M, Connelly MC, Rao C, Tibbe AG, Uhr JW, Terstappen LW. Tumor cells circulate in the peripheral blood of all major carcinomas but not in healthy subjects or patients with nonmalignant diseases. Clin Cancer Res 2004;10:6897-904.
10. Pinzani P, Salvadori B, Simi L, Bianchi S, Distante V, Cataliotti L, Pazzagli M, Orlando C. Isolation by size of epithelial tumor cells in peripheral blood of patients with breast cancer: correlation with real-time reverse transcriptase-polymerase chain reaction results and feasibility of molecular analysis by laser microdissection. Hum Pathol 2006;37:711-8.
11. Wong NS, Kahn HJ, Zhang L, Oldfield S, Yang LY, Marks A, Trudeau ME. Prognostic significance of circulating tumor cells enumerated after filtration enrichment in early and metastatic breast cancer patients. Breast Cancer Res Treat 2006;99:63-9.
12. Nagrath S, Sequist LV, Maheswaran S, Bell DW, Irimia D, Ulkus L, Smith MR, Kwak EL, Digumarthy S, Muzikansky A, Ryan P, Balis UJ, Tompkins RG, Haber DA, Toner M. Isolation of rare circulating tumor cells in cancer patients by microchip technology. Nature 2007;450:1235-9.
13. Bidard FC, Belin L, Delaloge S, Lerebours F, Ngo C, Reyal F, Alran S, Giacchetti S, Marty M, Lebofsky R, Poerga JY. Time-dependent prognostic impact of circulating tumor cells detection in non-metastatic breast cancer: 70-month analysis of the REMAGUS02 study. Int J Breast Cancer 2013;2013:130470.
14. Bidard FC, Proudhon C, Pierga JY. Circulating tumor cells in breast cancer. Mol Oncol 2016;10:419-30.
15. Lianidou ES, Markou A. Circulating tumor cells in breast cancer: detection systems, molecular characterization, and future challenges. Clin Chem 2011;57:1242-55.
16. Peeters DJ, De Laere B, Van den Eynden GG, Van Laere SJ, Rothé F, Ignatiadis M, Sieuwerts AM, Lambrechts D, Rutten A, van Dam PA, Pauwels P, Peeters M, Vermeulen PB, Dirix LY. Semiautomated isolation and molecular characterization of single or highly purified tumor cells from CellSearch enriched blood samples using dielectrophoretic cell sorting. Br J Cancer 2013;108:1358-67.
17. Smerage JB, Barlow WE, Hortobagyi GN, Winer EP, Leyland-Jones B, Srkalovic G, Tejwani S, Schott AF, O'Rourke MA, Lew DL, Doyle GV, Gralow JR, Livingston RB, Hayes DF. Circulating tumor cells and response to chemotherapy in metastatic breast cancer: SWOG S0500. J Clin Oncol 2014;32:3483-9.
18. Rack B, Schindlbeck C, Jückstock J, Andergassen U, Hepp P, Zwingers T, Friedl TW, Lorenz R, Tesch H, Fasching PA, Fehm T, Schneeweiss A, Lichtenegger W, Beckmann MW, Friese K, Pantel K, Janni W; SUCCESS Study Group. Circulating tumor cells predict survival in early average-to-high risk breast cancer patients. J Natl Cancer Inst 2014;106.
19. Jacob K, Sollier C, Jabado N. Circulating tumor cells: detection, molecular profiling and future prospects. Expert Rev Proteomics 2007;4:741-56.
20. Croset M, Goehrig D, Frackowiak A, Bonnelye E, Ansieau S, Puisieux A, Clézardin P. TWIST1 expression in breast cancer cells facilitates bone metastasis formation. J Bone Miner Res 2014;29:1886-99.
21. Zhou YM, Cao L, Li B, Zhang RX, Sui CJ, Yin ZF, Yang JM. Clinicopathological significance of ZEB1 protein in patients with hepatocellular carcinoma. Ann Surg Oncol 2012;19:1700-6.
22. Sanislo L, Vertakova-Krakovska B, Kuliffay P, Brtko J, Galbava A, Galbavy S. Detection of circulating tumor cells in metastatic breast cancer patients. Endocr Regul 2011;45:113-24.
23. Riethdorf S, Müller V, Zhang L, Rau T, Loibl S, Komor M, Roller M, Huober J, Fehm T, Schrader I, Hilfrich J, Holms F, Tesch H, Eidtman H, Untch M, von Minckwitz G, Pantel K. Detection and HER2 expression of circulating tumour cells: prospective monitoring in breast cancer patients treated in the neoadjuvant GeparQuattro trial. Clin Cancer Res 2010;16:2634-45.
24. Bogenrieder T, Herlyn M. Axis of evil: molecular mechanisms of cancer metastasis. Oncogene 2003;22:6524-36.
25. Potdar PD, Lotey NK. Role of circulating tumor cells in future diagnosis and therapy of cancer. J Cancer Metastasis Treat 2015;1:44-56.
26. Pantel K, Alix-Panabières C. Functional studies on viable circulating tumor cells. Clin Chem 2016;62:328-34.
27. Cayrefourcq L, Mazard T, Joosse S, Solassol J, Ramos J, Assenat E, Schumacher U, Costes V, Maudelonde T, Pantel K, Alix-Panabières C. Establishment and characterization of a cell line from human circulating colon cancer cells. Cancer Res 2015;75:892-901.
28. Yu M, Bardia A, Aceto N, Bersani F, Madden MW, Donaldson MC, Desai R, Zhu H, Comaills V, Zheng Z, Wittner BS, Stojanov P, Brachtel E, Sgroi D, Kapur R, Shioda T, Ting DT, Ramaswamy S, Getz G, Iafrate AJ, Benes C, Toner M, Maheswaran S, Haber DA. Cancertherapy. Ex vivo culture of circulating breast tumor cells for individualized testing of drug susceptibility. Science 2014;345:216-20.
29. Baccelli I, Schneeweiss A, Riethdorf S, Steiniger A, Vogel V, Klein C, Saini M, Bäuerle T, Wallwiener M, Holland-Letz T, Höfner T, Sprick M, Scharpff M, Marmé F, Sinn HP, Pantel K, Weichert W, Trumpp A. Identification of a population of blood circulating tumor cells from breast cancer patients that initiates metastasis in a xenograft assay. Nat Biotechnol 2013;31:539-44.
30. Gertler R, Rosenberg R, Fuehrer K, Dahm M, Nekarda H, Siewert JR. Detection of circulating tumor cells in blood using an optimized density gradient centrifugation. Recent Results Cancer Res 2003;162:149-55.
31. Gascoyne RC, Shim S. Isolation of circulating tumor cells by dielectrophoresis. Cancers (Basel) 2014;6:545-79.
32. Ashworth TR. A case of cancer in which cells similar to those in the tumors were seen in the blood after death. Aus Med J 1869;14:146-9.
33. Yu M, Stott S, Toner M, Maheswaran S, Haber DA. Circulating tumor cells: approaches to isolation and characterization. J Cell Biol 2011;192:373-82.
34. Cristofanilli M. Circulating tumor cells, disease progression, and survival in metastatic breast cancer. Semin Oncol 2006;33:S9-14.
35. Miller MC, Doyle GV, Terstappen LW. Significance of circulating tumor cells detected by the CellSearch system in patients with metastatic breast colorectal and prostate cancer. J Oncol 2010;2010:617421.
36. Mori S, Chang JT, Andrechek ER, Matsumura N, Baba T, Yao G, Kim JW, Gatza M, Murphy S, Nevins JR. Anchorage-independent cell growth signature identifies tumors with metastatic potential. Oncogene 2009;28:2796-805.
37. Uitto VJ, Schwartz D, Veis A. Degradation of basement-membrane collagen by neutral proteases from human leukocytes. Eur J Biochem 1980;105:409-17.
38. Venter JC, Adams MD, Myers EW, Li PW, Mural RJ, Sutton GG, Smith HO, Yandell M, Evans CA, Holt RA, Gocayne JD, Amanatides P, Ballew RM, Huson DH, Wortman JR, Zhang Q, Kodira CD, Zheng XH, Chen L, Skupski M, Subramanian G, Thomas PD, Zhang J, Gabor Miklos GL, Nelson C, Broder S, Clark AG, Nadeau J, McKusick VA, Zinder N, Levine AJ, Roberts RJ, Simon M, Slayman C, Hunkapiller M, Bolanos R, Delcher A, Dew I, Fasulo D, Flanigan M, Florea L, Halpern A, Hannenhalli S, Kravitz S, Levy S, Mobarry C, Reinert K, Remington K, Abu-Threideh J, Beasley E, Biddick K, Bonazzi V, Brandon R, Cargill M, Chandramouliswaran I, Charlab R, Chaturvedi K, Deng Z, Di Francesco V, Dunn P, Eilbeck K, Evangelista C, Gabrielian AE, Gan W, Ge W, Gong F, Gu Z, Guan P, Heiman TJ, Higgins ME, Ji RR, Ke Z, Ketchum KA, Lai Z, Lei Y, Li Z, Li J, Liang Y, Lin X, Lu F, Merkulov GV, Milshina N, Moore HM, Naik AK, Narayan VA, Neelam B, Nusskern D, Rusch DB, Salzberg S, Shao W, Shue B, Sun J, Wang Z, Wang A, Wang X, Wang J, Wei M, Wides R, Xiao C, Yan C, Yao A, Ye J, Zhan M, Zhang W, Zhang H, Zhao Q, Zheng L, Zhong F, Zhong W, Zhu S, Zhao S, Gilbert D, Baumhueter S, Spier G, Carter C, Cravchik A, Woodage T, Ali F, An H, Awe A, Baldwin D, Baden H, Barnstead M, Barrow I, Beeson K, Busam D, Carver A, Center A, Cheng ML, Curry L, Danaher S, Davenport L, Desilets R, Dietz S, Dodson K, Doup L, Ferriera S, Garg N, Gluecksmann A, Hart B, Haynes J, Haynes C, Heiner C, Hladun S, Hostin D, Houck J, Howland T, Ibegwam C, Johnson J, Kalush F, Kline L, Koduru S, Love A, Mann F, May D, McCawley S, McIntosh T, McMullen I, Moy M, Moy L, Murphy B, Nelson K, Pfannkoch C, Pratts E, Puri V, Qureshi H, Reardon M, Rodriguez R, Rogers YH, Romblad D, Ruhfel B, Scott R, Sitter C, Smallwood M, Stewart E, Strong R, Suh E, Thomas R, Tint NN, Tse S, Vech C, Wang G, Wetter J, Williams S, Williams M, Windsor S, Winn-Deen E, Wolfe K, Zaveri J, Zaveri K, Abril JF, Guigó R, Campbell MJ, Sjolander KV, Karlak B, Kejariwal A, Mi H, Lazareva B, Hatton T, Narechania A, Diemer K, Muruganujan A, Guo N, Sato S, Bafna V, Istrail S, Lippert R, Schwartz R, Walenz B, Yooseph S, Allen D, Basu A, Baxendale J, Blick L, Caminha M, Carnes-Stine J, Caulk P, Chiang YH, Coyne M, Dahlke C, Mays A, Dombroski M, Donnelly M, Ely D, Esparham S, Fosler C, Gire H, Glanowski S, Glasser K, Glodek A, Gorokhov M, Graham K, Gropman B, Harris M, Heil J, Henderson S, Hoover J, Jennings D, Jordan C, Jordan J, Kasha J, Kagan L, Kraft C, Levitsky A, Lewis M, Liu X, Lopez J, Ma D, Majoros W, McDaniel J, Murphy S, Newman M, Nguyen T, Nguyen N, Nodell M, Pan S, Peck J, Peterson M, Rowe W, Sanders R, Scott J, Simpson M, Smith T, Sprague A, Stockwell T, Turner R, Venter E, Wang M, Wen M, Wu D, Wu M, Xia A, Zandieh A, Zhu X. The sequence of the human genome. Science 2001;291:1304-51.
39. Chimal-Ramírez GK, Espinoza-Sánchez NA, Utrera-Barillas D, Benítez-Bribiesca L, Velázquez JR, Arriaga-Pizano LA, Monroy-García A, Reyes-Maldonado E, Domínguez-López ML, Pi-a-Sánchez P, Fuentes-Pananá EM. MMP1, MMP9, and COX2 expressions in promonocytes are induced by breast cancer cells and correlate with collagen degradation, transformation-like morphological changes in MCF-10A acini, and tumor aggressiveness. Biomed Res Int 2013;2013:279505.
40. Bozzuto G, Ruggieri P, Molinari A. Molecular aspects of tumor cell migration and invasion. Ann Ist Super Sanita 2010;46:66-80.
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Potdar PD, Sen K. Profiling of circulating tumor cells in liquid biopsies from metastatic cancer patients. J Cancer Metastasis Treat 2017;3:6-15. http://dx.doi.org/10.20517/2394-4722.2016.53
AMA Style
Potdar PD, Sen K. Profiling of circulating tumor cells in liquid biopsies from metastatic cancer patients. Journal of Cancer Metastasis and Treatment. 2017; 3: 6-15. http://dx.doi.org/10.20517/2394-4722.2016.53
Chicago/Turabian Style
Potdar, Pravin D., Keerti Sen. 2017. "Profiling of circulating tumor cells in liquid biopsies from metastatic cancer patients" Journal of Cancer Metastasis and Treatment. 3: 6-15. http://dx.doi.org/10.20517/2394-4722.2016.53
ACS Style
Potdar, PD.; Sen K. Profiling of circulating tumor cells in liquid biopsies from metastatic cancer patients. J. Cancer. Metastasis. Treat. 2017, 3, 6-15. http://dx.doi.org/10.20517/2394-4722.2016.53
About This Article
Special Issue
Copyright
Author Biographies

Data & Comments
Data
 Cite This Article 3 clicks
Cite This Article 3 clicks













Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.