Cancer, circulating tumor cells, and metastasis: could protein-derived peptide fragments impede brain metastases?
Abstract
The majority of cancer deaths can be attributed to cancer cell metastases that migrate to distant target organs. Brain metastases constitute one of the leading causes of morbidity and mortality among cancer patients, occurring in about 40% of patients with metastatic disease. Thus, there exists an unmet need for early detection, diagnosis, and treatment directed against early stage cancer cell metastasis. Previous studies have reported the development of methods to detect and identify early circulating tumor cells (CTCs) in the bloodstream prior to their seeding into distant organs. Using a comprehensive analysis of total CTCs mRNA content, investigators have developed a mRNA “transcriptome signature” of 126 genes involved in CTC metastatic events. The genes were parsed into various metastatic-related activities indicating that CTCs sustained a semi-dormancy state bent on: (1) stress survival; (2) metabolic maintenance; (3) DNA and translational stability; and (4) chemotactic pro-inflammatory capabilities. These activities suggested that CTCs might be susceptible to interactions with protein-derived peptide segments whose actions are involved with metastatic activities such as cell invasiveness, contact, adhesion, motility, spreading, and migration. The use of protein-derived (encrypted) peptides to impede CTC metabolic activities and disrupt signaling pathways could have therapeutic potential in patients with early metastatic disease.
Keywords
Introduction
Cancer metastases to the brain have been reported to occur in 10% to 20% of adult patients with malignant disease[1,2]. As such, brain metastases are one of the leading causes of morbidity and mortality among cancer patients with metastatic disease in the United States[3]. Indeed, the vast majority of cancer deaths can be attributed to tumor cell metastasis to distant sites rather than demise from the primary tumor mass itself. Several past reports have demonstrated that the presence of early circulating tumor cells (CTCs) provide a “feeder source” of break-away cells from the tumor mass cells that intravasate into the bloodstream to circulate, aggregate, then eventually migrate to metastatic sites of chemo-attracted target organs[4-6]. It may take extended time periods for the circulating cancer cells to develop aggregates detectable by radio-imaging as metastatic cell masses[7]. The early stages of disseminated tumor cells are able to aggregate to form micro-metastatic cell islets which further progress into macro-metastatic cell clusters. By the time the metastatic cells have migrated and “nested” into target tissues such as bone marrow and brain, the cells are already proliferating at exponential rates[8,9]. This advanced growth state of the tumor cell greatly reduces therapeutic options for the cancer patient. Therefore, it becomes crucial to detect and identify CTCs that represent very early stages of cells which disseminate from the primary tumor mass.
A recent report has, in fact, described use of a cutting-edge mRNA technology to characterize such CTCs. Hence, the first objective of the present commentary was to discuss the development of a biomarker mRNA signature that could screen and identify CTCs utilizing human breast cancer as the model. A second objective was to propose use of a potential therapeutic tool which could be directed against CTCs. These novel agents are referred to as “protein-derived peptide fragments” as discussed below.
Background studies
It may be deduced from the above discussion that early screening and identification of CTCs associated with metastasis could be beneficial for evaluation of treatment options and their responses to brain metastasis. In a recent study using breast cancer (BC) cells, Boral et al.[10] reasoned that mRNA characterization of CTCs from BC derived cells could provide a means for early diagnosis of brain metastases; this knowledge could aid in planning therapeutic strategies and determining the effectiveness of targeting metastatic cells. Boral’s studies confirmed that patients with BC produce CTCs that express high levels of biomarkers that are associated with regulation of cell growth, proliferation, and signal transduction pathways directly involved with metastasis. Using a comprehensive analysis of CTC total mRNA transcriptomes, these investigators derived a unique “transcriptome signature” (TS) that distinguished circulating BC cells from those of the primary tumor mass. Even more intriguing, the derived TS revealed distinct signaling pathways inherent to BC-derived circulating cells that could provide the means of metastases to the brain. The Boral’s report concluded that the CTC biomarker profile and knowledge of their signaling pathways could be valuable as screening tools for: (1) micro- and macro-metastatic cell characterization; (2) decision-making in treatment modalities; and (3) monitoring post-treatment responses in patients with metastatic disease.
Previous literature reports have emerged which enumerated epithelial cell adhesion molecules (EpCAM) present on CTCs, thus providing an estimate of the overall metastatic cell burden present in BC patients[11-13]. Using previous EpCAM-related studies as a starting point, Boral et al.[10] devised a computer-based genomic and mRNA workflow strategy which revealed a higher cell surface frequency and expression of metastatic-associated biomarkers in BC patient’s cells versus cells from healthy blood donors[10]. Finally, these investigators utilized parametric flow cytometry and MRI-proven metastasis brain scans to analyze their BC patient’s cell populations.
Components of the CTC signature
The CTC signature derived from circulating BC cells was parsed down to 126 genes involved in metastatic cell activities and signaling cascades[10]. Overall results from the 126 genes demonstrated that mRNA from 73 genes in the BC-CTC group were up-regulated while 53 genes were down-regulated. All of the CTC genes detected were distinct and none clustered with their corresponding cells and tissues from the primary BC mass. Some of the biomarker constituents of the CTC mRNA signature included cell activities involving growth regulation, cell adhesion, cell-to-cell contact, spreading, migration, and motility. Such biomarkers included CD86, PARP6, ERα, GBP2, Adam-17, DDIT4, SLC2A3, SRGN, and NOTCH-1. Additional biomarkers were involved with chemotaxis, pro-inflammatory factors and immunomodulatory networks which included CD44, CD45, CD24, TNF, IL-1B, NFkB, CXCL8, CXCR4, and PDGF-BB. Brain-related biomarkers encompassed NCAM, Serpin I1, plasmin, neuroserpin B2, and UPAR which are required for stealth transpassage through the blood brain barrier [Supplementary Table 1]. It is of interest that proteins such as plasmin, serpins, and UPAR are especially crucial to CTCs for brain entry[14].
A circumspect examination of some of the gene constituents of the CTC signature revealed that proteins related to various cellular activities and pathways could be parsed into several functional sub-groups. These groups displayed mRNA transcripts that were either enhanced (up-regulated) or reduced (down-regulated) in the blood circulating cells. The regulated gene transcripts encompassed cell activities such as: (1) growth and proliferation; (2) DNA transcription and translation; (3) signal transduction; (4) cell invasiveness and migration; and (5) mitotic and metabolic events. In summation, one could deduce from the above listing that CTC’s appeared to be groomed for maintaining a metabolic “status quo” semi-dormancy state in order to survive in the blood circulation while preparing for migration to a distant organ site[4,15]. While so doing, the CTC have to retain their functional cell maintenance in order to detach from cell-to-cell contacts, adhere to blood platelets, and migrate to target organs (i.e., bone marrow or brain) with the aid of inflammatory chemokine molecules.
Protein-encrypted peptides, CTCs, and metastasis
The containment of a class of growth factor, extra-cellular matrix, and angiogenic peptide fragments encrypted within the polypeptide chain of a full-length protein is known but is not widely recognized. However, some of the most potent growth inhibitors are derived from short peptide fragments (segments) already existent in naturally-occurring mammalian full length proteins that themselves affect cell growth and proliferation in an opposite function from the mother proteins. This less-recognized concept of a protein-derived reserve containing peptide growth Inhibitor fragments is becoming a recurring theme in the field of growth regulation, intracellular signaling, and cross-talk between signal transduction pathways. Classical examples of such occult (cryptic) peptides include the following examples; (1) tenacin binding peptide derived from fibronectin[16]; (2) angiostatin from plasmin[17]; (3) endostatin from type XVIII collagen[18]; (4) vasostatin from calreticulin[19]; and (5) constatin from type-IV collagen[20]. Such cryptic peptide sites can be exposed following a conformational change on a protein or can be released following proteolytic cleavage from a larger protein. These peptides can also be chemically synthesized as single fragments of 20-45 amino acids. A well-published example of a peptide site revealed following a conformational transition change on a full-length protein is an encrypted “growth inhibitory” site on alpha-fetoprotein (AFP), normally a growth promoting molecule[21-23]. The encrypted peptide segment, termed the growth inhibitory peptide (GIP), is a 34 amino acid segment concealed in a hydrophobic cleft of the completely-folded AFP molecule. The GIP site is revealed following protein unfolding in chemical environments containing high ligand concentrations of estrogens, fatty acids, and growth factors. This transitory GIP form converts the usually growth-enhancing AFP molecule into a growth-inhibiting polypeptide. This conversion occurs via protein un-folding into a conformational change resembling the denatured intermediate state of a molten globular form (MGF) of protein[22]. Since the MGF of AFP is a transitory intermediate form, AFP can refold back to its native tertiary fold following excess ligand removal. Because the AFP-MGF form is unstable, the GIP segment itself has now been synthesized, purified, and characterized as a distinct 34-mer synthetic peptide segment[23]. The 34-mer GIP fragment can inhibit both growth factor and estrogen-induced growth in a concentration-dependent fashion in addition to blocking metastatic-associated activities[24,25].
AFP-derived peptides, CTCs, and metastasis
It is germane to the present commentary that full-length AFP mRNA detected in CTCs from hepatocellular carcinoma patients has been reported to serve as a predictive marker for metastasis[26]. Furthermore, a computer bioinformatics study of multiple metastatic protein interactions with AFP (and its derived peptides) has recently been reported[27]. In that study, many of the "in silico" AFP interaction with metastatic associated proteins were experimentally confirmed. Both in vitro and in vivo BC studies have been performed using GIP which demonstrated both anti-growth and anti-metastatic activities. For example, in a microarray study, GIP was found to down-regulate the mRNA (1.5 to 8 fold) of many proteins detected in the “CTC signature” of the BC-derived circulating cells described by Boral et al.[10]. Such proteins included CD44, CD40, TNF, NFkB, IL-1 receptor, Serpin I-1, and p53 AIP1 among others. Many of these metastasis-associated proteins were reported to interact with AFP in protein-to-protein interactions; such metastases-related proteins included the laminin receptor, collagen-IV, Integrin B-1, IL-1B, and the neural cell adhesion molecule (NCAM). It is of interest that, Serpin-I1 and plasminogen activator are known to promote cancer cell survival in brain metastasis by means of brain plasmin inhibition[28]. In the pro-inflammatory arena, AFP itself was found to interact and block CCR5 and CXCR4 chemokine receptors which are required for metastatic BC cell migration[29-31].
Biological activities of AFP-derived peptides
Other sets of data involving AFP-derived peptides have been generated in vitro involving cancer cell adhesion, cell-to-cell interactions, cell spreading, motility, migration, and growth[32,33]. Regarding cancer cell proliferation, GIP was found to inhibit growth in multiple BC cell lines in vitro and to inhibit cell-to-cell contact inhibition overgrowth in cultured MCF-7 cells[33-35]. In addition, both full length AFP and GIP were both found capable of inhibiting platelet aggregation[36], a process necessary for CTC survival in the bloodstream; this activity involved integrins α2β1, α5β1, and α2β3. CTCs are known to adhere to blood vessel inner walls and to platelets, thereby cloaking themselves from circulating cytotoxic lymphocyte destruction[13,35,36]. Furthermore, GIP was found to block both adhesion of extra-cellular matrix (ECM) proteins to substrata as well as tumor cell adhesion to ECM-coated wells of microtiter plates[35]. The ECM proteins included laminin, fibrinogen, collagen-IV, fibronectin, thrombospondin, and vitronectin. In addition, both collagen-IV and NCAM have been reported to bind to the third domain of AFP at amino acid segments #433 to 545[34]. It has been further demonstrated in vitro that GIP notably interrupted the migration and invasion of follicular thyroid cancer cells[37]. It has also been reported that GIP could inhibit 60% of the cell spreading and migration of MCF-7 tumor cells in culture assays[34]. Because integrins and ECM proteins are both involved in cell migration by modulating the fine balance between cell-to-contact, adhesion, and cell detachment, it was noteworthy that GIP was found capable of disrupting the interaction between receptors and binding proteins in such activities. The final involvement of GIP with cancer cell activities was demonstrated using in vivo models of human BC xenografts in mice. GIP was reported to suppress cancer growth/proliferation in both xenograft and homograft models of MCF-7, GI-101, MDA-MB-231, and 6WI-1 BC tumors in host mice[25,32]. In the human MDA-MB-231 BC in vivo mouse model, GIP injections resulted in a 3-fold reduction in BC metastasis to the lungs as compared to controls. In the 6WI-1 in vivo mouse homograft model, GIP suppressed BC cell migration, invasiveness, and adherence to surrounding cells and tissues. Thus, GIP injections not only demonstrated BC growth suppression but also reduced metastatic-associated events in BC cells such as cell adherence, invasiveness, and migration in addition to decreased metastatic cell accumulation in distant organs. Finally, GIP administered in vitro produced an inhibition of cell membrane-induced agglutination, and induced cell shape changes via enhanced microtubule polymerization[24,34,35].
Concluding statements
It can be concluded from the above discourse that peptide fragments derived from plasma, ECM, and angiogenic-associated proteins are capable of tumor growth inhibition and suppression of metastatic spread. Such physiological events include cell-to-contact, cell migration, adhesion, detachment, spreading, and chemokine and receptor interactions. Following cell detachment from the BC tumor mass, the disseminated tumor cells extravasate through the tissue extracellular compartments, pass through disrupted (proteolysis) basement membranes, and emerge into the bloodstream. Once in the blood circulation, tumor cells can adhere and cluster into micro- and macro-metastatic islets that attach to blood platelets cloaking them from detection by cytotoxic lymphocytes. It is just prior to the stage of islet cluster formation that the metastatic cells are most vulnerable to blockade of signal transduction pathways [Figure 1]. Discovery of the CTC mRNA signature of CTCs en route to “nesting” in distant target organs, such as the brain, might allow investigators to design therapeutic strategies to impede metastatic invasion to the distant tissues and organs. One such group of potential metastatic disruptive agents could include plasma-, ECM-, and angiogenic protein-encrypted peptide fragments as discussed above. Such metastatic (migration) interfering peptides might be therapeutically beneficial to BC patients in early stages of micro-metastasis. Small peptides are known to have short half-lives (hours), little or no side effects, and could be intravenously administered. The screening of CTCs using the “signature” identification methodology, followed by peptide therapy, could potentially provide a novel 2-step detection/therapy strategy for select cancer patients with early metastatic disease.
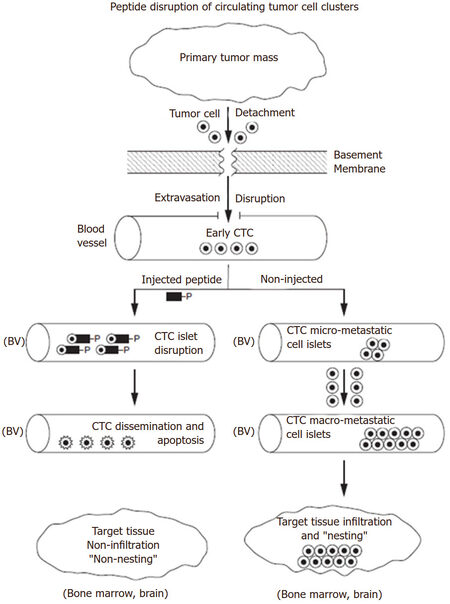
Figure 1. An injected peptide disruption of blood circulating tumor cells (CTCs) is depicted in the flow diagram. The primary mass of malignant tumors are known to shed cells which can migrate through the intercellular spaces en route to metastasis. The detached migratory tumor cells can extravasate through the blood vessel basement membrane and endothelial cell lining into the lumen of blood vessels (BV). Early CTCs soon form micro-metastatic clusters which further aggregate to form macro-metastatic islets (right side of diagram). CTCs eventually infiltrate into distal target tissues (bone marrow, brain) and “nest” there. However, if designer peptides home toward and bind to tumor cell membrane proteins/receptors as decoy ligands (see text for mRNA expressed proteins), tumor cell clusters could be disrupted and disseminated (left side of diagram). Single circulating cells including CTCs demonstrating cell membrane ruffling and disruption can become susceptible to apoptosis and/or immune surveillance destruction
Declarations
Authors’ contributionsMizejewski GJ contributed solely to the commentary.
Availability of data and materialsNot applicable.
Financial support and sponsorshipNone.
Conflicts of interestThe author declared that there are no conflicts of interest.
Ethical approval and consent to participateNot applicable.
Consent for publicationNot applicable.
Copyright© The Author(s) 2018.
REFERENCES
2. Stewart BW, Wild C, editors. World Cancer Report 2014. Lyon, France: International Agency for Research on Cancer; 2014. pp. 1-20.
3. Cooper JB, Ronecker JS, Tobias ME, Mohan AL, Hillard V, Murali R, Gandhi CD, Schmidt MH, Jhanwar-Uniyal M. Molecular sequence of events and signaling pathways in cerebral metastases. Anticancer Res 2018;38:1859-77.
4. Rack B, Schindlbeck C, Juckstock J, Andergassen U, Hepp P, Zwingers T, Friedl TW, Lorenz R, Tesch H, Fasching PA, Fehm T, Schneeweiss A, Lichtenegger W, Beckmann MW, Friese K, Pantel K, Janni W. Circulating tumor cells predict survival in early average-to-high risk breast cancer patients. J Natl Cancer Inst 2014;106:1-30.
5. Rhim AD, Mirek ET, Aiello NM, Maitra A, Bailey JM, McAllister F, Reichert M, Beatty GL, Rustgi AK, Vonderheide RH, Leach SD, Stanger BZ. EMT and dissemination precede pancreatic tumor formation. Cell 2012;148:349-61.
6. Meng S, Tripathy D, Frenkel EP, Shete S, Naftalis EZ, Huth JF, Beitsch PD, Leitch M, Hoover S, Euhus D, Haley B, Morrison L, Fleming TP, Herlyn D, Terstappen LW, Fehm T, Tucker TF, Lane N, Wang J, Uhr JW. Circulating tumor cells in patients with breast cancer dormancy. Clin Cancer Res 2004;10:8152-62.
8. Welch HG, Gorski DH, Albertsen PC. Trends in metastatic breast and prostate cancer--lessons in cancer dynamics. N Engl J Med 2015;373:1685-7.
10. Boral D, Vishnoi M, Liu HN, Yin W, Sprouse ML, Scamardo A, Hong DS, Tan TZ, Thiery JP, Chang JC, Marchetti D. Molecular characterization of breast cancer CTCs associated with brain metastasis. Nat Commun 2017;8:196-204.
11. Yu M, Bardia A, Wittner BS, Stott SL, Smas ME, Ting DT, Isakoff SJ, Ciciliano JC, Wells MN, Shah AM, Concannon KF, Donaldson MC, Sequist LV, Brachtel E, Sgroi D, Baselga J, Ramaswamy S, Toner M, Haber DA, Maheswaran S. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 2013;339:580-4.
12. Hyun KA, Koo GB, Han H, Sohn J, Choi W, Kim SI, Jung HI, Kim YS. Epithelial-to-mesenchymal transition leads to loss of EpCAM and different physical properties in circulating tumor cells from metastatic breast cancer. Oncotarget 2016;7:24677-87.
13. Jordan NV, Bardia A, Wittner BS, Benes C, Ligorio M, Zheng Y, Yu M, Sundaresan TK, Licausi JA, Desai R, O'Keefe RM, Ebright RY, Boukhali M, Sil S, Onozato ML, Iafrate AJ, Kapur R, Sgroi D, Ting DT, Toner M, Ramaswamy S, Haas W, Maheswaran S, Haber DA. HER2 expression identifies dynamic functional states within circulating breast cancer cells. Nature 2016;537:102-6.
14. Valiente M, Obenauf AC, Jin X, Chen Q, Zhang XH, Lee DJ, Chaft JE, Kris MG, Huse JT, Brogi E, Massague J. Serpins promote cancer cell survival and vascular co-option in brain metastasis. Cell 2014;156:1002-16.
15. Alix-Panabieres C, Pantel K. Clinical applications of circulating tumor cells and circulating tumor DNA as liquid biopsy. Cancer Discov 2016;6:479-91.
16. Ingham KC, Brew SA, Erickson HP. Localization of a cryptic binding site for tenascin on fibronectin. J Biol Chem 2004;279:28132-5.
17. Cao Y, Ji RW, Davidson D, Schaller J, Marti D, Sohndel S, McCance SG, O'Reilly MS, Llinas M, Folkman J. Kringle domains of human angiostatin. Characterization of the anti-proliferative activity on endothelial cells. J Biol Chem 1996;271:29461-7.
18. Folkman J. Antiangiogenesis in cancer therapy--endostatin and its mechanisms of action. Exp Cell Res 2006;312:594-607.
19. Pike SE, Yao L, Jones KD, Cherney B, Appella E, Sakaguchi K, Nakhasi H, Teruya-Feldstein J, Wirth P, Gupta G, Tosato G. Vasostatin, a calreticulin fragment, inhibits angiogenesis and suppresses tumor growth. J Exp Med 1998;188:2349-56.
20. Kamphaus GD, Colorado PC, Panka DJ, Hopfer H, Ramchandran R, Torre A, Maeshima Y, Mier JW, Sukhatme VP, Kalluri R. Canstatin, a novel matrix-derived inhibitor of angiogenesis and tumor growth. J Biol Chem 2000;275:1209-15.
21. Mizejewski GJ, MacColl R. Alpha-fetoprotein growth inhibitory peptides: potential leads for cancer therapeutics. Mol Cancer Ther 2003;2:1243-55.
22. Uversky VN, Kirkitadze MD, Narizhneva NV, Potekhin SA, Tomashevski A. Structural properties of alpha-fetoprotein from human cord serum: the protein molecule at low pH possesses all the properties of the molten globule. FEBS Lett 1995;364:165-7.
23. Mizejewski GJ, Dias JA, Hauer CR, Henrikson KP, Gierthy J. Alpha-fetoprotein derived synthetic peptides: assay of an estrogen-modifying regulatory segment. Mol Cell Endocrinol 1996;118:15-23.
24. Mizejewski G, Smith G, Butterstein G. Review and proposed action of alpha-fetoprotein growth inhibitory peptides as estrogen and cytoskeletal-associated factors. Cell Biol Int 2004;28:913-33.
25. Mizejewski GJ, Butterstein G. Survey of functional activities of alpha-fetoprotein derived growth inhibitory peptides: review and prospects. Curr Protein Pept Sci 2006;7:73-100.
26. Jin J, Niu X, Zou L, Li L, Li S, Han J, Zhang P, Song J, Xiao F. AFP mRNA level in enriched circulating tumor cells from hepatocellular carcinoma patient blood samples is a pivotal predictive marker for metastasis. Cancer Lett 2016;378:33-7.
27. Mizejewski GJ. Third domain ligand binding fragment of alpha-fetoprotein: detection of metastasis-associated molecular targets. Cancer Therapy Oncol Int J 2017;6:1-15.
28. Valiente M, Obenauf AC, Jin X, Chen Q, Zhang XH, Lee DJ, Chaft JE, Kris MG, Huse JT, Brogi E, Massague J. Serpins promote cancer cell survival and vascular co-option in brain metastasis. Cell 2014;156:1002-16.
29. Mizejewski GJ. Mechanism of cancer growth suppression of alpha-fetoprotein derived growth inhibitory peptides (GIP): comparison of GIP-34 versus GIP-8 (AFPep). Updates and prospects. Cancers (Basel) 2011;3:2709-33.
30. Atemezem A, Mbemba E, Marfaing R, Vaysse J, Pontet M, Saffar L, Charnaux N, Gattegno L. Human alpha-fetoprotein binds to primary macrophages. Biochem Biophys Res Commun 2002;296:507-14.
31. Mizejewski GJ. Alpha-fetoprotein as a biomarker for immunodeficiency diseases: relevance to ataxia telangiectasia & related disorders. J Immunodefic Disor 2014;3:1-12.
32. Mizejewski GJ, Muehlemann M, Dauphinee M. Update of alpha fetoprotein growth-inhibitory peptides as biotherapeutic agents for tumor growth and metastasis. Chemotherapy 2006;52:83-90.
33. Mizejewski GJ. Role of integrins in cancer: survey of expression patterns. Proc Soc Exp Biol Med 1999;222:124-38.
34. Muehlemann M, Miller KD, Dauphinee M, Mizejewski GJ. Review of growth inhibitory peptide as a biotherapeutic agent for tumor growth, adhesion, and metastasis. Cancer Metastasis Rev 2005;24:441-67.
35. Mizejewski GJ, Mirowski M, Garnuszek P, Maurin M, Cohen BD, Poiesz BJ, Posypanova GA, Makarov VA, Severin ES, Severin SE. Targeted delivery of anti-cancer growth inhibitory peptides derived from human alpha-fetoprotein: review of an International Multi-Center Collaborative Study. J Drug Target 2010;18:575-88.
36. Czokalo M, Tomasiak M. Alpha fetoprotein inhibits aggregation of human platelets. Haematologia (Budap) 1989;22:11-8.
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Mizejewski GJ. Cancer, circulating tumor cells, and metastasis: could protein-derived peptide fragments impede brain metastases?. J Cancer Metastasis Treat 2018;4:27. http://dx.doi.org/10.20517/2394-4722.2018.20
AMA Style
Mizejewski GJ. Cancer, circulating tumor cells, and metastasis: could protein-derived peptide fragments impede brain metastases?. Journal of Cancer Metastasis and Treatment. 2018; 4: 27. http://dx.doi.org/10.20517/2394-4722.2018.20
Chicago/Turabian Style
Mizejewski, Gerald J.. 2018. "Cancer, circulating tumor cells, and metastasis: could protein-derived peptide fragments impede brain metastases?" Journal of Cancer Metastasis and Treatment. 4: 27. http://dx.doi.org/10.20517/2394-4722.2018.20
ACS Style
Mizejewski, GJ. Cancer, circulating tumor cells, and metastasis: could protein-derived peptide fragments impede brain metastases?. J. Cancer. Metastasis. Treat. 2018, 4, 27. http://dx.doi.org/10.20517/2394-4722.2018.20
About This Article
Copyright
Data & Comments
Data
 Cite This Article 3 clicks
Cite This Article 3 clicks












Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.