Cryptic NUP214-ABL1 fusion with complex karyotype, episomes and intra-tumor genetic heterogeneity in a T-cell lymphoblastic lymphoma
Abstract
T-lymphoblastic lymphoma (T-LBL) is a rare and aggressive form of non-Hodgkin’s lymphoma and little is known about their molecular background. However, complex karyotypes were already related to this group of malignancy and associated with poor outcome. Here, we describe a 17-year-old female being diagnosed with T-LBL and a normal karyotype after standard G-banding with trypsin-Giemsa (GTG)-banding. However, further analyses including high-resolution molecular approaches, array-comparative genomic hybridization (aCGH), multiplex ligation-dependent probe amplification, fluorescence in situ hybridization and multicolor chromosome banding revealed a cryptic complex karyotype, NUP214-ABL1 gene fusion, episomes and intra-tumor genetic heterogeneity. In addition, homozygous loss of CDKN2A, as well as amplification of oncogene TLX1 (HOX11) were detected. Actually, NUP214-ABL1 fusion gene replicated autonomously in this case as episomes. Overall, highly amplification of NUP214-ABL1 fusion gene defines possibly a new subgroup of T-LBL patients which accordingly could benefit from treatment with tyrosine kinase inhibitors. As episomes are missed in standard karyotyping aCGH should be performed routinely in T-LBL to possibly detect more of such cases.
Keywords
Introduction
Lymphoblastic lymphoma (LBL) is a rare and aggressive form of non-Hodgkin’s lymphoma (NHL). LBL develops from immature B cells committed to the B- (B-LBL) or T-cell lineage (T-LBL). LBL is morphologically indistinguishable from acute lymphoblastic leukemia (ALL) and 90% of it have a T-cell phenotype. LBL also accounts for approximately 2% of all NHL cases and occur in adult, children and adolescent, with a male predominance (three time more male are affected)[1-2].
Chromosomal abnormalities in T-LBL are not well defined and cytogenetic data in T-LBL is limited. However, a few published cytogenetic studies revealed that typical chromosomal aberrations identified in T-cell ALL (T-ALL) are also present in T-LBL. These include translocations of T-cell receptor (TCR) gene to genes encoding transcription factors such as TAL1, TLX1, LMO2, and LYL1. In particular, the translocation t(9;17)(q34;q22~23) is typically found in T-LBL[1-4]. However, no single recurrent and typical genetic alteration for T-LBL could be identified. This is in contrast to other malignancies like translocation of ALK gene in anaplastic large cell lymphoma, MYC gene in Burkitt lymphoma or BCL2 gene in follicular lymphoma.
Here we present the comprehensive analysis of a T-LBL case with a normal karyotype, according to standard G-banding with trypsin-Giemsa (GTG)-banding, using high resolution molecular methods, identifying also some intra-tumor genetic heterogeneity besides unusual acquired genetic alterations. Also here we report NUP214-ABL1 gene fusion in this patient, which appears cryptic due to its localization in episomes.
Case report
A seventeen-year-old female patient, who was initially diagnosed in South Africa with T-ALL, presented in the clinic in Poland with abdominal pain, accompanied by diarrhea and vomiting; she was here initially treated only symptomatically. A few days before, a blood test already revealed hyperleukocytosis (589 × 109/l) with presence of 94% lymphoblasts in blood smear, hemoglobin 8.5 g/dl, and platelet count 53 × 109/l. Bone marrow findings were: hypercellularity with 95% lymphoblasts, lack of megakaryocytes and Periodic-Acid-Schiff (PAS) staining identified in 70% of the blasts thick grains (data not shown). Ultrasound of abdomen showed enlargement of the spleen to 152 mm, and presence of fluid in the lower pelvis. Cervical lymph nodes were bilaterally enlarged with diameters of 3-4 cm, and small submandibular nodes were bilaterally enlarged to 2 cm in diameter.
Cytogenetic and immunophenotypic analyses were done. The latter characterized a T-LBL due to high expression of CD45 (100%), CD2 (96.6%), CD4 (97.3%), CD8 (90%), CD7 (77.1%), CD5 (76.0%), sCD3 (71.2%), CD1a (70.0%) and the lack of TdT, CD19, CD34 and CD38.
Banding cytogenetic analyses were done in unstimulated bone marrow cells according to standard procedures[5] from the material taken at initial diagnoses. A total of 20 metaphases were available and analyzed on a banding resolution of 300 bands per haploid karyotype, revealing a normal female karyotype. Molecular diagnostic polymerase chain reaction (PCR)-based tests for presence of gene fusions BCR/ABL (p190 and p210), TCF3/PBX1, MLL/AF4 and SIL/TAL1 were negative (results not shown).
Also genomic DNA isolated from cells fixed in acetic acid-methanol (1:3) was subjected to array-comparative genomic hybridization (aCGH) as well as the multiplex ligation probe amplification (MLPA) studies, as previously reported[6], Finally, fluorescence in situ hybridization (FISH) was done[6-8], revealing a highly complex karyotype [Figure 1 and Table 1] with gene-amplification due to episomes (abbreviated here as epi), which can be reported as:
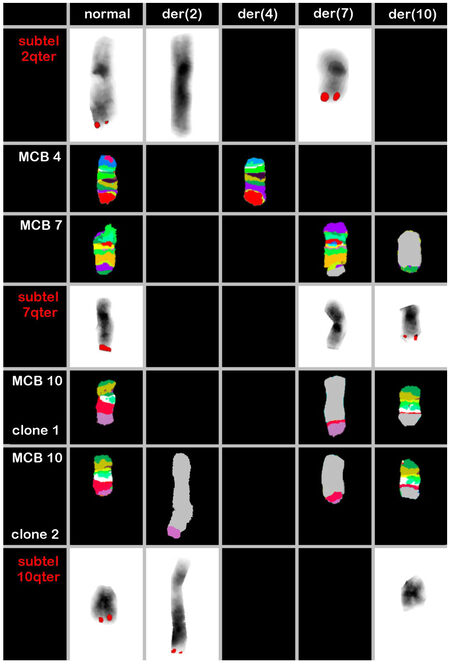
Figure 1. Result of multicolor banding (MCB) probesets for chromosomes 4, 7 and 10 are shown. MCB 10 showed the founder clone and subclone. Locus-specific probes (LSPs) for chromosomes 2, 7 and 10 characterized the breakpoints in 2q37.3, 7p34, 10q24.3 and 10q25.1 [Table 1]. The final karyotype after application of all approaches is summarized in the text. der = derivative chromosome
Locus specific probes used for FISH together with their location according to genome browser version NCBI/hg18; this version was used here as some here applied FISH-probes are no longer available in newer genome browser versions. Results obtained are presented using standard (gene) abbreviations and such used according to the international system of cytogenomic nomenclature
| Cytoband | Position [NCBI36/hg18] | Genes/locus | Probe | Result (signals on…) |
|---|---|---|---|---|
| 2q37.1 | chr2:234,552,641-234,701,765 | n.d | RP11-263G22 | der(2) |
| 2q37.2 | chr2:236,163,266-236,349,539 | n.d. | RP11-473L20 | der(2) |
| 2q37.3 | chr2:238,251,662-238,463,936 | n.d. | RP11-497D24 | der(7) |
| 2q37.3 | chr2:242,433,475-242,633,697 | D2S447 | 2qTEL (Vysis) | der(7) |
| 6q23.3 | chr6:135,544,146-135,582,003 | MYB | SPEC MYB DCBAP (Zytovision) | amp(6)(q23.3q23.3) |
| 7q31.2 | chr7:116,099,695-116,225,676 | MET | SPEC MET/CEN7 (Zytovision) | der(7) |
| 7q33 | chr7:133,287,726-133,474,337 | n.d. | RP11-639H21 | der(7) |
| 7q33 | chr7:134,684,542-134,842,811 | n.d. | RP11-371N6 | der(7) |
| 7q33 | chr7:136,263,935-136,416,924 | n.d. | RP11-88K4 | der(7) |
| 7q33 | chr7:137,919,273-138,093,873 | n.d. | RP11-269N18 | der(7) |
| 7q34 | chr7:141,674,679-141,819,906 | TCRB | n.a. | n.a. |
| 7q34 | chr7:142,124,883-142,316,809 | n.d. | RP11-39H3 | der(10) |
| 7q34-q35 | chr7:142,787,852-142,859,896 | n.d. | RP11-811J9 | der(10) |
| 7q35 | chr7:143,536,879-143,690,749 | n.d. | RP11-45N9 | der(10) |
| 7q35 | chr7:145,715,880-145,867,471 | n.d. | RP11-97H18 | der(10) |
| 7q35 | chr7:147,084,270-147,259,380 | n.d. | RP11-302C22 | der(10) |
| 7q36.3 | chr7:158,400,001-158,600,424 | VIJyRM2000 | 7qTEL (Vysis) | der(10) |
| 9p21.3 | chr9:21,792,635-21,984,490 | MTAP CDKN2A/B | SPEC CDKN2A/CEN9 (Zytovision) | del(9)(p21.3p21.3) |
| 9q34.13 | chr9:132,579,089-132,752,883 | ABL1 | LSI BCR, ABL (Vysis) | amp(9)(q34.12q34.12) |
| 10q23.31 | chr10:89,613,175-89,718,512 | PTEN | SPEC PTEN/CEN10 (Zytovision) | der(10) |
| 10q24.31 | chr10:102,880,252-102,887,526 | TLX1 | n.a. | n.a. |
| 10q24.31-q32 | chr10:102,895,115-103,074,760 | n.d. | RP11-324L3 | der(7) |
| 10q24.32 | chr10:104,652,453-104,813,482 | n.d. | RP11-724N1 | der(7) |
| 10q25.1 | chr10:106,748,189-106,912,787 | n.d. | RP11-165P9 | der(7) |
| 10q25.1 | chr10:107,741,530-107,812,754 | n.d. | RP11-596L14 | der(7) |
| 10q25.2 | chr10:112,350,581-112,499,609 | n.d. | RP11-364E8 | der(2) |
| 10q25.2 | chr10:116,774,286-116,971,219 | n.d. | RP11-338L11 | der(2) |
| 10q26.13 | chr10:123,227,834-123,347,962 | FGFR2 | SPEC FGFR2/CEN10 (Zytovision) | der(2) |
| 10q26.3 | chr10:134,925,980-135,126,361 | D10S2290 | 10qTEL (Vysis) | der(2) |
46,XX,der(2)t(2;7)(q37.3;q25.1),del(4)(p14p16),t(7;10)(q34;q24),del(9)(p21.3p21.3),epi(6;9)(q23.3;q34.12)x20~30[20%]/46,XX,der(2)t(2;7)(q37.3;q25.1),del(4)(p14p16),der(7)(7pter->7q34::10q24.1->10q25.1::2q37.3->2qter),del(9)(p21.3p21.3),der(10)t(10;7)(q23;q34),epi(6;9)(q23.3;q34.12)x20~30[40%]/46,XX[40%].
In the FISH-studies done here, between 15 and 25 metaphases were evaluated per applied probe-set, thus in the final karyotype overall percentages are given for the observed clones.
NUP214-ABL1 fusion could be deduced from aCGH data - the region being amplified ends on one side at NUP214- and on other side at ABL1-gene - as the amplified region is present as episomes, which are circular, there must be NUP214-ABL1 fusion.
The patient was treated according to the Polish Adult Leukemia Group (PALG) protocol, with induction therapy consisting of prednisone, daunorubicin, vincristine and PEG-L-asparaginase. No remission was achieved and the patient was re-treated according to fludarabine, cytarabine, and mitoxantrone (FLAM) with consolidation course (metrotrexate, cyclophosphamide and PEG-L-asparaginase) and maintenance treatment. After ten months, the patient relapsed and was now treated according to Hyper-CVAD protocol (cyclophosphamide, vincristine, doxorubicin/Adriamycin and dexamethasone). Still, one month later the patient unfortunately died.
Discussion
Recurrent acquired genetic lesions play a key role in predicting and assessing risks, so are the treatment protocols to be applied. Still, little is known about the copy number alterations (CNAs) accompanying structural abnormalities in T-LBL, such as the NUP214-ABL1 fusion gene. ABL1 fusion proteins are sensitive to tyrosine kinase inhibitors, which potentially can be included in future treatment strategy and NUP214 is a component of the nuclear pore complex, which mediates nucleocytoplasmic transport. NUP214 is widely expressed and is involved in the pathogenesis of acute myeloid leukaemia associated with the translocation t(6;9)(p23;q34) as DEK-NUP214 fusion[9-11].
To the best of our knowledge, a cryptic NUP214-ABL1 fusion yet has only been identified in 6% of individuals with T-ALL and is the second most prevalent fusion gene involving ABL1[12-15]. Here we report this for the first time in a T-LBL case, and even detected it as a high level amplification; most probably after inversion, duplication or translocation, gene fusion, circularization and amplification happened. As ABL1 is one of the best targetable tyrosine kinases, identification of ABL1 gene fusion is clinically important, as patients may potentially benefit from tyrosine kinase inhibitors[16-17].
Episomes are submicroscopic, circular and large acentric DNA fragments that can replicate autonomously. One of the common formation-mechanisms for extrachromosomal elements in cancer cells is episome-replication and unequal segregation during cell division, resulting finally in an increase of copy numbers. Still they that are invisible in banding cytogenetics; this is because episomes are composed of only several hundred kilobases of amplified oncogenes and/or drug-resistance genes, and thus are too small to be visualized by light-microscopy[18-20]. Of interest, we detected a variable number of episomes (20-30) in different cells. However, we suggest that c-MYB is also present on the same episomes [Figure 2]; due to lack of material we could not confirm this by FISH.
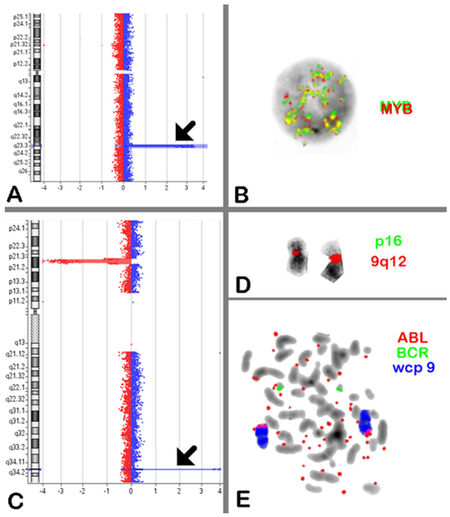
Figure 2. A: Array comparative genomic hybridization (aCGH) analysis of chromosome 6 revealed high level of 6q23.3 amplification containing MYB gene (arrow); B: MYB Dual Color Break Apart Probe was applied and showed high level of amplification more than 20 copies/per cell; C: aCGH analysis of chromosome 9 revealed biallelic deletion of CDKN2A at 9p21.3 and high level of 9q34 amplification contains ABL1 and NUP214 (arrow); D: FISH confirmed the homozygous deletion of CDKN2A in metaphase; E: BCR, ABL Dual Color Probe was applied and showed variable number of episomes (20-30) in spread metaphases. aCGH: array-comparative genomic hybridization; FISH: fluorescence in situ hybridization; wcp: whole chromosome paint
Additionally, recurrent acquired CNAs in different chromosomal regions were also identified besides unique ones for this case [Table 2]. Taken together, the genomic abnormalities in T-ALL and T-LBL are so similar that they could be considered as identical diseases in the future[1,4,12,14,15,21-24].
ummary of CNAs detected by aCGH. Recurrent (R) and unique (U) acquired CNAs are correspondingly highlighted in first column. Results obtained are presented using standard (gene) abbreviations and such used according to the international system of cytogenomic nomenclature
| Chromosome (alteration U or R) | Cytobands | GRCH37/hg19 | Size of imbalance [Mb] | Genes |
|---|---|---|---|---|
| 1 (U) | del(1)(p36.31p36.23) | chr1:5,958,728-7,238,618 | 1.27 | NPHP4, KCNAB2, CHD5, RPL22, RNF207, ICMT, HES3, GPR153, ACOT7, HES2, ESPN, MIR4252, TNFRSF25, PLEKHG5, NOL9, TAS1R1, ZBTB48, KLHL21, PHF13, THAP3, DNAJC11, CAMTA1 |
| del(1)(q22.2q22.2) | chr1:91,620,826-91,739,326 | 0.2 | HFM1 | |
| 4 (U) | del(4)(p16.3p14) | chr4:3,072,509-38,882,925 | 35.8 | HTT, C4orf44, RGS12, HGFAC, DOK7, LRPAP1, LOC100133461, ADRA2C, LOC348926, OTOP1, TMEM128, LYAR, ZBTB49, D4S234E, STX18, LOC100507266, MSX1, CYTL1, STK32B, C4orf6, EVC2, EVC, CRMP1, JAKMIP1, LOC285484, WFS1, PPP2R2C, MAN2B2, MRFAP1, LOC93622, S100P, MRFAP1L1, CNO, KIAA0232, TBC1D14, LOC100129931, CCDC96, TADA2B, GRPEL1, FLJ36777, SORCS2, PSAPL1, MIR4274, AFAP1 AS1, AFAP1, ABLIM2, SH3TC1, HTRA3, ACOX3, METTL19, GPR78, CPZ, HMX1, LOC650293, USP17, USP17L6P, DEFB131, MIR548I2, DRD5, SLC2A9, WDR1, MIR3138, ZNF518B, CLNK, MIR572, HS3ST1, HSP90AB2P, RAB28, LOC285547, NKX3-2, LOC285548, BOD1L, LOC152742, CPEB2, C1QTNF7, CC2D2A, FBXL5, FAM200B, BST1, CD38, FGFBP1, FGFBP2, PROM1, TAPT1, FLJ39653, LDB2, QDPR, CLRN2, LAP3, MED28, FAM184B, DCAF16, NCAPG, LCORL, SLIT2, LOC100505893, MIR218-1, PACRGL, KCNIP4, NCRNA00099, LOC100505912, GPR125,GBA3, PPARGC1A, MIR573, DHX15, SOD3, CCDC149, LGI2, SEPSECS, LOC285540, PI4K2B, ZCCHC4, ANAPC4, SLC34A2, SEL1L3, C4orf52, RBPJ, CCKAR, TBC1D19, STIM2, MIR4275, PCDH7, ARAP2, DTHD1, KIAA1239, C4orf19,RELL1, PGM2,TBC1D1, PTTG2,FLJ13197, KLF3,TLR10, TLR1,TLR6, FAM114A1,MIR574 |
| 6 (R) | amp(6)(q23.3q23.3) | chr6:134,245,761-136,118,354 | 1.87 | TBPL1, SLC2A12, HMGA1P7, SGK1, ALDH8A1, HBS1L, MIR3662, MYB, AHI1, NCRNA00271 |
| 9 (R) | del(9)(p21.3p21.3) | chr9:20,605,923-21,218,606 | 0.61 | MLLT3, KIAA1797, MIR491, PTPLAD2, IFNB1, IFNW1, IFNA21,IFNA4, IFNA7, IFNA10, IFNA16 |
| del(9)(p21.3p21.3) | chr9:21,252,517-23,002,377 | 1.75 | IFNA22P, IFNA5, KLHL9, IFNA6, IFNA13, IFNA2, IFNA8, IFNA1, LOC554202, IFNE, MIR31, MTAP, C9orf53, CDKN2A, CDKN2B-AS1, CDKN2B, DMRTA1 | |
| amp(9)(q34.1q34.1) | chr9:133,658,293-134,092,544 | 0.43 | ABL1, QRFP, FIBCD1, LAMC3, AIF1L, NUP214 | |
| 10 (U) | del(10)(q25.1q25.2) | chr10:111,634,169-112,348,580 | 0.71 | XPNPEP1, ADD3, MXI1, SMNDC1, DUSP5, SMC3 |
| 15 (U) | amp(15)(q13.3q13.3) | chr15:32,098,670-32,539,666 | 0.44 | CHRNA7 |
As shown in our case, NUP214-ABL1 is accompanied with loss of cyclin-dependent kinase inhibitor 2A (CDKN2A), which encodes the tumorsuppressors p16INK4A and p14ARF, and affects cell cycle progression. CDKN2A gene deletion can be detected at initial diagnosis or acquired at relapse, suggesting that CDKN2A gene deletion is a secondary genetic event and associated with chromosomal rearrangements. This may as a result lead to the aberrant expression of a diverse group of T-cell-specific transcription factors, which again can function as oncogenes, such as TLX1 and TLX3[2,21,25]. The translocation t(7;10)(q34;q24), resulting from the TRB/TLX1 fusion gene, has been reported in several studies, and is present in 5% of pediatric and 30% of adult with T-cell ALL[25-28].
Overall, in the present T-LBL case we identified substantial intra-tumor genetic heterogeneity and complexity. The founder clone has TRB/TLX1 fusion gene and the subclone has TRB/TLX1 fusion gene plus complex karyotype involving three-way translocation t(2;7;10)(2q37.3;7q34;10q25.1), further developing into a more complex subclone. Interestingly, the breakpoints at 2q37.3, 7q34, 10q24.3 and 10q25.1 were not previously reported in T-LBL[29]. Thus, this data provides genetic support for a multi-step pathogenesis: deletion of a tumor-suppressor gene (CDKN2A), deregulated expression of a transcription factor TLX1 and most likely overexpression of a constitutively activated tyrosine kinase (NUP214-ABL1) and oncogene c-MYB due to episome amplification and the unique phenotypes of the T-LBL case mentioned above.
To conclude, this study demonstrates the power of high resolution molecular approaches. It may be considered that the use of such approaches is the most efficient and future standard method for screening ABL1 alteration. Particularly in T-LBL patients this may be advantageous, as ABL1 modulates T-cell development and plays a role in cytoskeletal remodeling processes in T-cells. Besides, the intra-tumor genetic heterogeneity in cancer has important implications for reservoirs of cells involved in progression of disease and drug resistance therapy. As NUP214-xABL1 fusion is sensitive to the tyrosine kinase inhibitor, this suggests that new therapeutic approaches in T-LBL may improve outcome and/or decrease treatment-related morbidity.
Declarations
Authors’ contributionsDid the FISH-studies and drafted the paper: Othman MAK
Performed array comparative genomic hybridization (aCGH) analyses and interpretation: Melo JB, Carreira IM, Othman MAK
Provided T-LBL-case including clinical and banding cytogenetic data: Grygalewicz B, Kołkowska-Leśniak A
Planned and organized the study and did final drafting of the paper: Liehr T
All authors read and approved the paper.
Availability of data and materialsAll data is provided in this article. Also the patient was mentioned previously in[30] as P61.
Financial support and sponsorshipNone.
Conflicts of interestAll authors declared that there are no conflicts of interest.
Ethical approval and consent to participateThe present study was approved by the Ethical Board at the Friedrich Schiller University (Jena, Germany; approval No., 1105‑04/03). Consent to participate of the parent of the patient studied here is available on request from the authors of this paper.
Consent for publicationNot applicable.
Copyright© The Author(s) 2018.
REFERENCES
1. Sekimizu M, Sunami S, Nakazawa A, Hayashi Y, Okimoto Y, Saito AM, Horibe K, Tsurusawa M, Mori T. Chromosome abnormalities in advanced stage T-cell lymphoblastic lymphoma of children and adolescents: a report from Japanese Paediatric Leukaemia/Lymphoma Study Group (JPLSG) and review of the literature. Br J Haematol 2011;154:612-7.
2. Baleydier F, Decouvelaere AV, Bergeron J, Gaulard P, Canioni D, Bertrand Y, Lepretre S, Petit B, Dombret H, Beldjord K, Molina T, Asnafi V, Macintyre E. T cell receptor genotyping and HOXA/TLX1 expression define three T lymphoblastic lymphoma subsets which might affect clinical outcome. Clin Cancer Res 2008;14:692-700.
3. Lones MA, Heerema NA, Le Beau MM, Sposto R, Perkins SL, Kadin ME, Kjeldsberg CR, Meadows A, Siegel S, Buckley J, Abromowitch M, Kersey J, Bergeron S, Cairo MS, Sanger WG. Chromosome abnormalities in advanced stage lymphoblastic lymphoma of children and adolescents: a report from CCG-E08. Cancer Genet Cytogenet 2007;172:1-11.
4. Ellison DA, Parham DM, Sawyer JR. Cytogenetic findings in pediatric T-lymphoblastic lymphomas: one institution's experience and a review of the literature. Pediatr Dev Pathol 2005;8:550-6.
5. Claussen U, Michel S, Mühlig P, Westermann M, Grummt UW, Kromeyer-Hauschild K, Liehr T. Demystifying chromosome preparation and the implications for the concept of chromosome condensation during mitosis. Cytogenet Genome Res 2002;98:136-46.
6. Othman MA, Grygalewicz B, Pienkowska-Grela B, Rygier J, Ejduk A, Rincic M, Melo JB, Carreira IM, Meyer B, Liehr T. A novel IGH@ gene rearrangement associated with
7. Weise A, Mrasek K, Fickelscher I, Claussen U, Cheung SW, Cai WW, Liehr T, Kosyakova N. Molecular definition of high-resolution multicolor banding probes: first within the human DNA sequence anchored FISH banding probe set. J Histochem Cytochem 2008;56:487-93.
8. Liehr T, Claussen U. Current developments in human molecular cytogenetic techniques. Curr Mol Med 2002;2:283-97.
9. Kang ZJ, Liu YF, Xu LZ, Long ZJ, Huang D, Yang Y, Liu B, Feng JX, Pan YJ, Yan JS, Liu Q. The Philadelphia chromosome in leukemogenesis. Chin J Cancer 2016;35:48.
11. Hagemeijer A, Graux C. ABL1 rearrangements in T-cell acute lymphoblastic leukemia. Genes Chromosomes Cancer 2010;49:299-308.
12. Graux C, Cools J, Melotte C, Quentmeier H, Ferrando A, Levine R, Vermeesch JR, Stul M, Dutta B, Boeckx N, Bosly A, Heimann P, Uyttebroeck A, Mentens N, Somers R, MacLeod RA, Drexler HG, Look AT, Gilliland DG, Michaux L, Vandenberghe P, Wlodarska I, Marynen P, Hagemeijer A. Fusion of NUP214 to ABL1 on amplified episomes in T-cell acute lymphoblastic leukemia. Nat Genet 2004;36:1084-9.
13. Chiaretti S, Tavolaro S, Ghia EM, Ariola C, Matteucci C, Elia L, Maggio R, Messina M, Ricciardi MR, Vitale A, Ritz J, Mecucci C, Guarini A, Foà R. Characterization of ABL1 expression in adult T-cell acute lymphoblastic leukemia by oligonucleotide array analysis. Haematologica 2007;92:619-26.
14. Graux C, Stevens-Kroef M, Lafage M, Dastugue N, Harrison CJ, Mugneret F, Bahloula K, Struski S, Grégoire MJ, Nadal N, Lippert E, Taviaux S, Simons A, Kuiper RP, Moorman AV, Barber K, Bosly A, Michaux L, Vandenberghe P, Lahortiga I, De Keersmaecker K, Wlodarska I, Cools J, Hagemeijer A, Poirel HA; Groupe Francophone de Cytogénétique Hématologique,. Heterogeneous patterns of amplification of the NUP214-ABL1 fusiongene in T-cell acute lymphoblastic leukemia. Leukemia 2009;23:125-33.
15. Basso K, Mussolin L, Lettieri A, Brahmachary M, Lim WK, Califano A, Basso G, Biondi A, Cazzaniga G, Rosolen A. T-cell lymphoblastic lymphoma shows differences and similarities with T-cell acute lymphoblastic leukemia by genomic and gene expression analyses. Genes Chromosomes Cancer 2011;50:1063-75.
16. Clarke S, O'Reilly J, Romeo G, Cooney J. NUP214-ABL1 positive T-cell acute lymphoblastic leukemia patient shows an initial favorable response to imatinib therapy post relapse. Leuk Res 2011;35:e131-3.
17. Duployez N, Grzych G, Ducourneau B, Alarcon Fuentes M, Grardel N, Boyer T, Abou Chahla W, Bruno B, Nelken B, Clappier E, Preudhomme C. NUP214-ABL1 fusion defines a rare subtype of B-cell precursor acute lymphoblastic leukemia that could benefit from tyrosine kinase inhibitors. Haematologica 2016;101:e133-4.
18. Maurer BJ, Lai E, Hamkalo BA, Hood L, Attardi G. Novel submicroscopic extrachromosomal elements containing amplified genes in human cells. Nature 1987;327:434-7.
20. Turner KM, Deshpande V, Beyter D, Koga T, Rusert J, Lee C, Li B, Arden K, Ren B, Nathanson DA, Kornblum HI, Taylor MD, Kaushal S, Cavenee WK, Wechsler-Reya R, Furnari FB, Vandenberg SR, Rao PN, Wahl GM, Bafna V, Mischel PS. Extrachromosomal oncogene amplification drives tumour evolution and genetic heterogeneity. Nature 2017;543:122-5.
21. Kim HJ, Woo HY, Koo HH, Tak EY, Kim SH. ABL oncogene amplification with p16(INK4a) gene deletion in precursor T-cell acute lymphoblastic leukemia/lymphoma: report of the first case. Am J Hematol 2004;76:360-3.
22. Stergianou K, Fox C, Russell NH. Fusion of NUP214 to ABL1 on amplified episomes in T-ALL--implications for treatment. Leukemia 2005;19:1680-1.
23. Kawamata N, Zhang L, Ogawa S, Nannya Y, Dashti A, Lu D, Lim S, Schreck R, Koeffler HP. Double minute chromosomes containing MYB gene and NUP214-ABL1 fusion gene in T-cell leukemia detected by single nucleotide polymorphism DNA microarray and fluorescence in situ hybridization. Leuk Res 2009;33:569-71.
24. De Keersmaecker K, Porcu M, Cox L, Girardi T, Vandepoel R, de Beeck JO, Gielen O, Mentens N, Bennett KL, Hantschel O. NUP214-ABL1-mediated cell proliferation in T-cell acute lymphoblastic leukemia is dependent on the LCK kinase and various interacting proteins. Haematologica 2014;99:85-93.
25. Ferrando AA, Neuberg DS, Dodge RK, Paietta E, Larson RA, Wiernik PH, Rowe JM, Caligiuri MA, Bloomfield CD, Look AT. Prognostic importance of TLX1 (HOX11) oncogene expression in adults with T-cell acute lymphoblastic leukaemia. Lancet 2004;363:535-6.
26. Cauwelier B, Dastugue N, Cools J, Poppe B, Herens C, De Paepe A, Hagemeijer A, Speleman F. Molecular cytogenetic study of 126 unselected T-ALL cases reveals high incidence of TCRbeta locus rearrangements and putative new T-cell oncogenes. Leukemia 2006;20:1238-44.
27. Le Noir S, Ben Abdelali R, Lelorch M, Bergeron J, Sungalee S, Payet-Bornet D, Villarèse P, Petit A, Callens C, Lhermitte L, Baranger L, Radford-Weiss I, Grégoire MJ, Dombret H, Ifrah N, Spicuglia S, Romana S, Soulier J, Nadel B, Macintyre E, Asnafi V. Extensive molecular mapping of TCRα/δ- and TCRβ-involved chromosomal translocations reveals distinct mechanisms of oncogene activation in T-ALL. Blood 2012;120:3298-309.
28. Liu Y, Easton J, Shao Y, Maciaszek J, Wang Z, Wilkinson MR, McCastlain K, Edmonson M, Pounds SB, Shi L, Zhou X, Ma X, Sioson E, Li Y, Rusch M, Gupta P, Pei D, Cheng C, Smith MA, Auvil JG, Gerhard DS, Relling MV, Winick NJ, Carroll AJ, Heerema NA, Raetz E, Devidas M, Willman CL, Harvey RC, Carroll WL, Dunsmore KP, Winter SS, Wood BL, Sorrentino BP, Downing JR, Loh ML, Hunger SP, Zhang J, Mullighan CG. The genomic landscape of pediatric and young adult T-lineage acute lymphoblastic leukemia. Nat Genet 2017;49:1211-8.
29. Cancer Genome Anatomy Project. Mitelman database of chromosome aberrations and gene fusions in cancer. Available from: http://cgap.nci.nih.gov/Chromosomes/Mitelman. [Last accessed on 14 Aug 2018].
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Othman MAK, Grygalewicz B, Kołkowska-Leśniak A, Melo JB, Carreira IM, Liehr T. Cryptic NUP214-ABL1 fusion with complex karyotype, episomes and intra-tumor genetic heterogeneity in a T-cell lymphoblastic lymphoma. J Cancer Metastasis Treat 2018;4:50. http://dx.doi.org/10.20517/2394-4722.2018.41
AMA Style
Othman MAK, Grygalewicz B, Kołkowska-Leśniak A, Melo JB, Carreira IM, Liehr T. Cryptic NUP214-ABL1 fusion with complex karyotype, episomes and intra-tumor genetic heterogeneity in a T-cell lymphoblastic lymphoma. Journal of Cancer Metastasis and Treatment. 2018; 4: 50. http://dx.doi.org/10.20517/2394-4722.2018.41
Chicago/Turabian Style
Othman, Moneeb A.K, Beate Grygalewicz, Agnieszka Kołkowska-Leśniak, Joana B. Melo, Isabel M. Carreira, Thomas Liehr. 2018. "Cryptic NUP214-ABL1 fusion with complex karyotype, episomes and intra-tumor genetic heterogeneity in a T-cell lymphoblastic lymphoma" Journal of Cancer Metastasis and Treatment. 4: 50. http://dx.doi.org/10.20517/2394-4722.2018.41
ACS Style
Othman, MAK.; Grygalewicz B.; Kołkowska-Leśniak A.; Melo JB.; Carreira IM.; Liehr T. Cryptic NUP214-ABL1 fusion with complex karyotype, episomes and intra-tumor genetic heterogeneity in a T-cell lymphoblastic lymphoma. J. Cancer. Metastasis. Treat. 2018, 4, 50. http://dx.doi.org/10.20517/2394-4722.2018.41
About This Article
Special Issue
Copyright
Data & Comments
Data

 Cite This Article 19 clicks
Cite This Article 19 clicks












Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.