The molecular interaction of ADAMTS-1 and fibulin-1 and its potential contribution to breast cancer biology
Abstract
Aim: Fibulins and ADAMTSs are two families of extracellular matrix proteins implicated in key functional and pathological processes. The fact that the fibulin-1 and ADAMTS-1 proteins interact raises new questions about the roles of these extracellular matrix proteins in modulating tumor progression. Herein, we described the functional implications of the interaction between fibulin-1 and ADAMTS-1 on the behavior of breast cancer cell lines.
Methods: Fibulin-1 and ADAMTS-1 were exogenously expressed in MCF-7 and MDA-MB-231 cell lines to assay the effect of their interaction in cellular properties.
Results: ADAMTS-1 expression exacerbates tumor effects in terms of proliferation, invasion and mammosphere formation. In contrast, the simultaneous expression of ADAMTS-1 and fibulin-1 impairs these effects. The analysis of the expression of both proteins in human breast cancer tissue arrays provides new insights into the complex roles of fibulin-1 and ADAMTS-1 in this type of tumor.
Conclusion: Our results suggests that the interaction between ADAMTS-1 and fibulin-1 induces a pronounced anti-tumoral effect.
Keywords
Introduction
The extracellular environment is a dynamic niche in which multiple molecular interactions occur that govern cell fate. Several families of extracellular proteins function in this molecular dance, influencing cell properties and thus regulating either physiological or pathological processes[1,2]. In addition, stromal cells are considered especially important in the extracellular environment as they induce extracellular modifications that can promote dysfunction in processes such as in cancer progression[3].
In general, proteases are considered to be responsible for altering tissues and initiating the extracellular remodeling of tissues. Among proteases, members of the ADAMTS family have been widely implicated in different steps of cancer development. The ADAMTS family consists of 19 members, all of which have several domains, including a metalloprotease, a disintegrin and a variable number of thrombospondin motifs, which allow them to interact with different components of the extracellular matrix (ECM)[4]. ADAMTS-1 was the first identified member of this family of matrix metalloproteases due to its association with inflammation processes[5]. Since its discovery, ADAMTS-1 has been described to participate in several other processes, such as organogenesis, vessel formation and ovulation[6-8]. The participation of ADAMTS-1 in cancer is underlined by its implication in some of the most important features of cancer development and progression[9]. Thus, ADAMTS-1 has positive effects on cell survival, invasion and migration processes and, in general, does not participate in cell proliferation[10,11]. Furthermore, ADAMTS-1 promotes angiogenesis in breast xenografts, but it is also able to inhibit angiogenesis in lung and hepatic metastasis as well as in other cell-based angiogenic experiments[11,12].
Several studies provide conflicting associations of ADAMTS-1 with breast cancer; ADAMTS-1 has been reported to be both a pro- and anti-tumorigenic factor[10,13-15]. This dual function of ADAMTS-1 depends on the conditions in which the ADAMTS-1 pathway is functioning, i.e., microRNAs such as miR-365 or peroxisome proliferator-activated receptor (PPARδ)[16,17]. The tumor promotion or tumor inhibition properties associated with ADAMTS-1 can also depend on the fragmentation of ADAMTS-1 or the interactions of ADAMTS-1 with other components of the ECM[18]. One of the known partners of ADAMTS-1 in the ECM is fibulin-1; the interaction of these two proteins was described as a result of a yeast two-hybrid screen for potential interactions of ADAMTS-1. As a result of this interaction, the proteolytic activity of ADAMTS-1 towards aggrecan is increased; thus, fibulin-1 is considered a cofactor of ADAMTS-1 aggrecanase activity[19].
Fibulin-1 was the first identified member of the fibulin family of matrix proteins[20]. The members of this family each contain a fibulin-like domain at the carboxy terminal of the protein. The fibulin family consists of seven members of different lengths that contain different motifs with varying functions, such as anaphylatoxin domains (fibulin-1 and fibulin-2), epidermal growth factor (EGF)-like modules (fibulin-3 and fibulin-4), thrombospondin-like and von Willebrand factor domains (fibulin-6) or a sushi domain (fibulin-7)[21]. In particular, fibulin-1 and fibulin-2 are closely related and characterized by the presence of anaphylatoxin and calcium-binding EGF (cbEGF) modules. However, only fibulin-2 contains two additional modules: cysteine (cys)-rich and cys-free domains. In addition, fibulin-1, fibulin-2, fibulin-3, fibulin-4 and fibulin-5 are considered elastogenic fibulins since they are important components of elastic fibers that are particularly abundant in the dermis, lungs and arterial walls[21,22].
The roles of fibulins in tumor development have been widely discussed, and they show both pro- and anti-tumor activities[23]. In particular, fibulin-1 has been described to be overexpressed in various human cancers and associated with processes such as invasion, migration, and in vivo tumor growth[24-26]. The role of fibulin-1 in breast cancer depends on multiple factors. For example, four alternative forms of fibulin-1 have been described (fibulin-1A, fibulin-1B, fibulin-1C and fibulin-1D), which differ at the carboxy terminal[27]. Fibulin-1D acts as a product of a tumor suppressor gene, whereas fibulin-1C (lacks only 21 amino acids of the fibulin-like domain) behaves as an oncogenic protein[25,28]. Furthermore, the identification of fibulin-1 localization or fibulin-1-expressing cells are also important metrics since fibulin-1 is one of the genes specifically expressed in normal fibroblasts but not in breast cancer-associated fibroblasts[29]. The epigenetic downregulation of fibulin-1 has also been described in various cancers, such as gastric cancer, colorectal cancer, hepatocellular carcinoma, and bladder cancer, but this downregulation has not been described in breast cancer[30-33].
Herein, we describe the effects of the ADAMTS-1/fibulin-1 interaction on breast cancer cell properties related to cancer progression and development. We found that cell proliferation, invasion and migration are susceptible to being affected by this interaction as well as the capacity of tumorsphere formation, which reflects the capability of cancer cells to form tumor focus. Finally, we detected very low levels of ADAMTS-1 expression and high levels of fibulin-1 expression in a human tissue array from breast cancer patients, which can be considered a deviation towards a cancerous phenotype.
Methods
Cells lines and cell culture conditions
Human MDA-MB-231 and murine MCF-7 breast cancer cell lines were kindly provided by Dr. Carlos López-Otín (University of Oviedo, Spain). Cells were routinely maintained in Dulbecco’s Modified Eagle’s medium (DMEM) containing 10% fetal bovine serum, 50 μg/mL streptomycin and 100 U/mL penicillin (Life Technologies). For transfection experiments, vectors containing the full-length human cDNAs for fibulin-1 (kindly provided by Dr. Tatako Sasaki, Oita University, Japan) and Flag-tagged ADAMTS-1 (Origene) were transfected into cells at 75% confluence using TransIT-X2 (Mirus) as recommended by the manufacturer. Cells transfected with an empty vector were used as a control. Stably expressing transfectants were selected with 500 μg/mL G418 (Sigma-Aldrich) and used for all the experiments.
Western blotting and immunoprecipitation
For western blot analysis, proteins were resolved by 10% polyacrylamide gel electrophoresis, transferred to a nitrocellulose membrane and subsequently probed with the indicated antibodies. Primary antibodies for the detection of ADAMTS-1 were purchased from Sigma-Aldrich (FLAG M2, for immunoprecipitation) and Bioss (bs-1208R-A488, for immunohistochemistry and western blotting), and those used for the detection of fibulin-1 were obtained from Novus (NBP1-84725, for immunohistochemistry) and Santa Cruz Biotechnology (H-190, for western blotting).
The anti-actin antibody used as the load control was purchased from Sigma-Aldrich (AC-15). Immunoreactive proteins were visualized using HRP-peroxidase-labeled anti-rabbit or anti-mouse secondary antibodies and an electrochemiluminescence (ECL) detection system (Pierce). For immunoprecipitation, MCF-7 double-transfected cells (ADAMTS-1 and fibulin-1) were resuspended in lysis buffer (100 mM Tris-HCl at pH 7.4, 150 mmol/L NaCl, 10 mmol/L EDTA, 1% desoxycholic acid, 1% Triton X-100, and 0.1% SDS containing one complete protease inhibitor cocktail (one tablet/50 mL buffer; Roche Molecular Biochemicals) and incubated for 2 h on ice with eventual stirring. Cell debris was removed by centrifugation in a bench-top centrifuge for 15 min at 4 ℃. Protein concentration was quantified using the BCA Protein Assay Kit from Pierce Technology. For immunoprecipitation, 1 mg of protein extract was incubated with anti-FLAG M2 affinity gel (Sigma-Aldrich) for 16 h at 4 ℃ following the manufacturer’s instructions. After three washes in lysis buffer, the immunoprecipitates were resolved by western blotting.
Invasion assays
In vitro invasion potential was evaluated using 24-well Matrigel-coated invasion chambers with an 8-μm pore size (BD Biosciences). For MCF-7 cells, 5 × 105 cells were allowed to migrate for 96 h using 10% fetal bovine serum as a chemoattractant. Cells that reached the lower surface were stained with crystal violet. At least three independent experiments were performed in triplicate for each condition. Cells were counted in eight randomly selected microscopic fields. In the case of MDA-MB-231 cells, invasion was evaluated after 24 h.
Migration assays
The migratory capacity of cells on the ECM components fibronectin, laminin I, and type-I collagen was examined using the Radius™ 24-Well Cell Migration Assay kit (Cell Biolabs) following the manufacturer’s instructions. Briefly, 5 × 105 cells were seeded per well, and migration was monitored by time-lapse microscopy using a Zeiss Axio Observer Microscopy. Experiments were performed in triplicate, and the covered area was quantified at different times using ImageJ. For MCF-7 cells, the results were obtained after 24 h of migration. In the case of MDA-MB-231 cells, we used a barrier-migration assay over a period of 24 h and the same ECM components[34].
Cell proliferation determination by Ki-67 staining
Cell proliferation was estimated by the quantification of Ki-67-positive nuclei in MCF-7 and MDA-MB-231 cell cultures, including controls and the different transfectants. Cells were fixed with 4% paraformaldehyde, blocked with 10% fetal bovine serum and incubated overnight with an anti-Ki67 antibody (Santa Cruz Biotechnologies). After three washes in phosphate buffered saline (PBS), the cells were incubated with a secondary Alexa 546-conjugated antibody (Life Technologies) for 1 h. In all samples, DAPI (100 ng/mL) was added to visualize the DNA in the cell nucleus. Images were obtained using a fluorescence microscope (Axiovert). After quantification, the data were plotted as an average of Ki-67-positive nuclei in relation to the total number of nuclei per microscopic field.
Mammosphere cultures
A total of 4 × 104 MCF-7 cells were plated in 6-well ultralow attachment plates (Costar) and grown in MammoCult Basal Medium (Stem Cell Research) supplemented with 10% MammoCult Proliferation Supplement, 4 μg/mL heparin and 0.5 μg/mL hydrocortisone. After 7 days, mammospheres were collected and enzymatically dissociated as previously described[35]. The individual dissociated cells were cultured in 96-well ultralow attachment plates at a density of 20 cells/well. Mammosphere formation was microscopically monitored daily to ensure that the mammospheres were derived from single cells and not from aggregates. The number of mammospheres was determined after 7 days of culture.
Human breast cancer tissue array
A breast cancer tissue array containing samples from different tumor stages was employed to evaluate ADAMTS1 and FBLN1 expression in human breast cancer samples. The tissue array was obtained from the Institute of Oncology of Asturias Tumour Bank. Written informed consent was obtained from all patients prior to sample collection. The study was approved by the appropriate institutional review board according to national and EU guidelines.
Slides were processed for indirect peroxidase immunohistochemistry as follows: sections were deparaffinized, rehydrated and then rinsed in PBS containing 1% Tween-20. For the detection of fibulin-1 and ADAMTS-1, the sections were heated in an Envision FLEX target retrieval solution at high pH and 80 ℃ for 20 min and then incubated for 20 min at room temperature in the same solution. Endogenous peroxidase activity (3% H2O2) and nonspecific binding (33% fetal calf serum) were blocked, and the sections were incubated overnight at 4 ℃ with primary antibodies diluted 1:100. We used labeled polymer-HRP (ready for use) antibodies from DAKO as the secondary antibodies. 3-3’-Diaminobenzidine was employed as a chromogen. Selected slides were counterstained with hematoxylin to ascertain structural details. As immunostaining controls, representative sections were processed in the same way as described above using nonimmune rabbit or mouse sera instead of the primary antibodies or by omitting the primary antibodies during the incubation. Furthermore, when available, additional controls were carried out using specifically preabsorbed antisera.
Fibulin-1 immunostaining intensity was semiquantitatively evaluated by two independent observers directly under the microscope using ten randomly selected fields (20x objective and a 20x ocular). The intensity of the immunoreaction was evaluated as strong (+++), moderate (++), weak (+) or negative (-).
Survival analysis
To assess the effect of ADAMTS-1 and fibulin-1 on breast cancer prognosis, survival probability was determined using the data available at www.kmplot.com[36]. The results are presented as Kaplan-Meier longrank test survival plots.
Statistical analysis
Statistical analysis was carried out using Microsoft Office software. Data are presented as mean ± SE. The occurrence of significant differences was determined with the Student-Welch t test. P values under 0.05 were considered statistically significant (*P < 0.05, **P < 0.01, ***P < 0.005).
Results
ADAMTS-1 and fibulin-1 interact in breast cancer cell lines
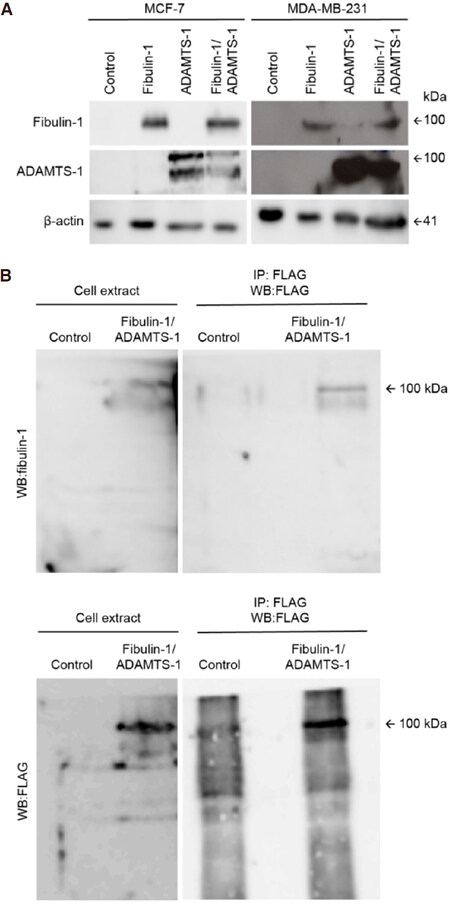
Fibulin-1 has been described as an ADAMTS-1 cofactor because of its proteolytic activity towards aggrecan degradation[19]. Fibulin-1 is not an ADAMTS-1 substrate; instead, it is an interacting partner that can affect cancer-associated properties such as the migration, invasion and proliferation of breast cancer cell lines. Thus, we used two common human breast cancer cell lines, MCF-7 and MDA-MB-231. First, we quantified the endogenous expression of both proteins in cell extracts obtained from both cell lines [Figure 1]. No detectable levels of fibulin-1 or ADAMTS-1 were visualized by western blotting in either the MCF-7 or the MDA-MB-231 cell extracts [Figure 1A]. Thus, we transfected both cell lines with the corresponding cDNAs (ADAMTS-1, fibulin-1 or both simultaneously) to obtain transfectants expressing one of the proteins or both proteins together. For control purposes, we used an empty vector as a negative control [Figure 1A, control]. Once we obtained all the transfectants, we tried to probe the in vivo interaction of ADAMTS-1 and fibulin-1 in MCF-7 extracts by immunoprecipitation. To this end, we took advantage of the Flag epitope present in the exogenously expressed ADAMTS-1 protein. Fibulin-1 western blotting following Flag (ADAMTS-1) immunoprecipitation showed a 100 kDa reactive band only in extracts from the ADAMTS-1/fibulin-1 double-transfected cells [Figure 1B, top], which demonstrated the existence of this interaction in the MCF-7 double transfectants [Figure 1B]. To probe the presence of ADAMTS-1 in the same immunoprecipitate, we carried out a Flag western blot that showed the presence of a 100 kDa reactive band that corresponded to the size of ADAMTS-1 [Figure 2, bottom].
Figure 1. Selection of MCF-7 and MDA-MB-231 stable transfectants and coimmunoprecipitation of fibulin-1 with ADAMTS-1. A: Western blot analysis of MCF-7 and MDA-MB-231 expressing exogenous fibulin-1, ADAMTS-1 or both proteins simultaneously (Fibulin-1/ADAMTS-1). Control, cells transfected with an empty vector. Actin was used as a loading control. Molecular weight marker is indicated on the right. Independent gels were performed for each western blot; B: immunoprecipitation of MCF-7 control cell extracts or extracts from cells expressing fibulin-1 and ADAMTS-1 (Fibulin-1/ADAMTS-1) using an anti-FLAG M2 affinity gel. IP, immunoprecipitated. Top, detection with an anti-fibulin-1 antibody. Bottom, detection with an anti-FLAG (ADAMTS-1) antibody. Intervening irrelevant lanes are not shown
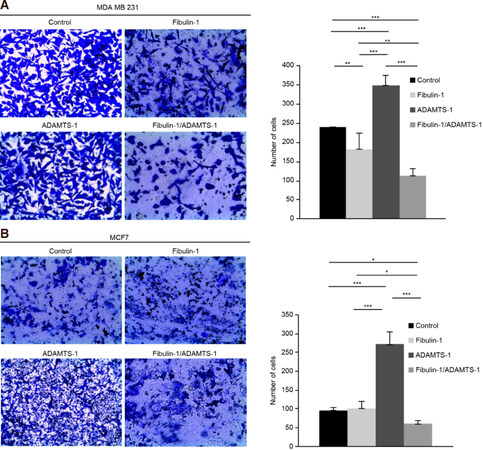
Figure 2. Coexpression of ADAMTS-1 and fibulin-1 reduces the invasion properties of the breast cancer cell lines MCF-7 and MDA-MB-231. Cell invasion assay using Matrigel-coated invasion chambers. A: representative microscopic pictures of MDA-MB-231 cells expressing exogenous fibulin-1, ADAMTS-1 or both proteins simultaneously. Cells transfected with an empty vector were used as a control. Cells that reached the lower surface were counted and are graphically shown; B: representative microscopic pictures of MCF-7 cells expressing exogenous fibulin-1, ADAMTS-1 or both proteins simultaneously. Cells transfected with an empty vector were used as a control. Cells that reached the lower surface were counted and graphically represented. P values under 0.05 were considered statistically significant (*P < 0.05, **P < 0.01, ***P < 0.005)
Effect of the fibulin-1/ADAMTS-1 interaction on invasion properties
After we demonstrated the fibulin-1/ADAMTS-1 interaction in breast cancer cells, we decided to explore the functional consequences of this event by first checking the effects of this interaction on the invasion properties of both cell lines. To address invasion properties, we used Matrigel-coated invasion chambers [Figure 2], and we observed that the fibulin-1/ADAMTS-1 interaction resulted in a significant reduction in the invasion capability of both cell lines. Thus, the invasive potential of the double transfectants of both the MDA-MB-231 and the MCF-7 cell lines was decreased compared with that of the control cell lines (MDA-MB-231: average of 112 double transfectants cells/field vs. 240 control cells/field; MCF-7: 65 double transfectants cells/field vs. 90 control cells/field). The expression of fibulin-1 alone had the ability to reduce the invasiveness of MDA-MB-231 cells (an average of 175 vs. 240 cells/field) but not the invasiveness of MCF-7 cells. In contrast, and in agreement with the previously described tumor-promoting properties of ADAMTS-1, the invasion capabilities of the ADAMTS-1 transfectants of both cell lines showed an important increase compared with those of the control cell lines (averages of 340 vs. 240 cells/field for the MDA-MB-231 cell line and 275 vs. 90 cells/field in MCF-7 cells).
Effect on the migration properties of the fibulin-1/ADAMTS-1 interaction
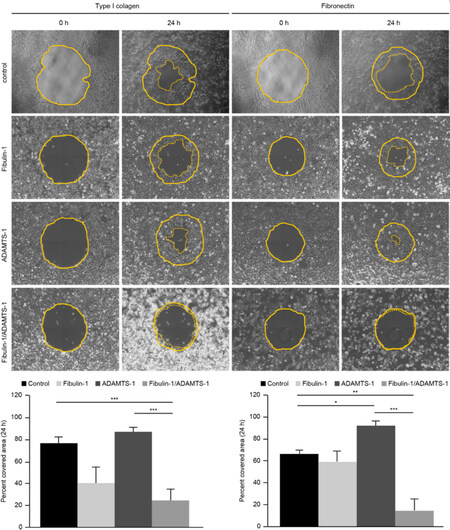
Next, we analyzed the migration potential of both cell lines over two components of the extracellular matrix, type-I collagen and fibronectin. After 24 h, we measured the covered area and graphed the results for a better interpretation of the results. In all cases, substrates and cell lines, we observed that the migration properties of the double transfectants (fibulin-1 and ADAMTS-1) were significantly compromised compared with those of their respective controls. We observed that the control MCF-7 cells were able to cover 79% of the area on type-1 collagen-coated dishes, whereas the double transfectants only were able to cover 25% of the area. In the case of fibronectin, the control cells could cover 65% of the dish area, and the double transfectants could cover 18% of the dish area [Figure 3]. The data for the control and double transfectants cells of the MDA-MB-231 cell line were somewhat similar; the control cells covered 90% of the areas of the type-1 collagen- and fibronectin-coated plates, while the double transfectants covered 35% and 25% of the type-1 collagen- and fibronectin-coated plates, respectively [Supplementary Figure 1]. The expression of ADAMTS-1 alone also showed significant effects on migration only over the fibronectin substrate compared with the control cells and double transfectants of the MCF-7 and MDA-MB-231 cell lines [Figure 3 and Supplementary Figure 1, respectively]. The overexpression of fibulin-1 alone showed significant effects compared with the double-transfected and ADAMTS-1-overexpressing MDA-MB-231 cells over fibronectin and type-1 collagen [Supplementary Figure 1].
Figure 3. Coexpression of ADAMTS-1 and fibulin-1 reduces the migration properties of the breast cancer cell line MCF-7 over type-I collagen and fibronectin. Top, Representative images of the migration of MCF-7 transfectants on wells coated with type-I collagen and fibronectin using the RadiusTM 24-Well Cell Migration Assay kit (Cell Biolabs). MCF-7 cells transfected with an empty vector were used as a control. Pictures of the initial (t: 0 h) and final (t: 24 h) times are included. The starting point is indicated with a thin dotted line, and the final point is indicated with a thick dotted line. Bottom, graphical representation of percentages of the area covered after 24 h (from three independent experiments)
Effects of the fibulin-1/ADAMTS-1 interaction on cellular proliferation
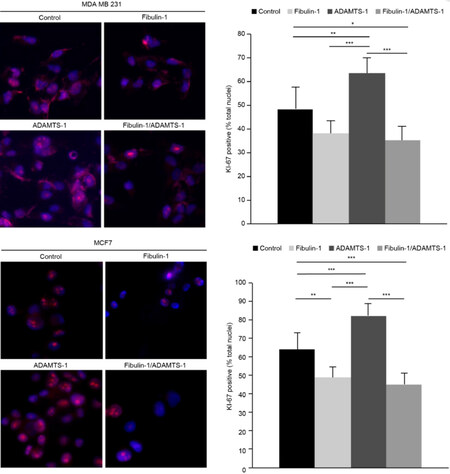
Ki-67 nuclear staining is a good marker to measure the proliferative state of any given cell line. In this study, we used Ki-67 nuclear staining to evaluate the proliferative properties of all the transfectants generated. In both cell lines, MDA-MB-231 and MCF-7, ADAMTS-1 overexpression increased cell proliferation, as shown in Figure 4. The number of Ki-67-positive nuclei in the ADAMTS-1-expressing and control cells of the MDA-MB-231 cell line was 63 and 48, respectively, whereas that in the ADAMTS-1-expressing and control cells of the MCF-7 cell line was 82 and 64, respectively. On the other hand, fibulin-1 overexpression was able to significantly reduce proliferation in only the MCF-7 cells; 49 and 64 Ki-67-positive nuclei were detected in the fibulin-overexpressing and control cells, respectively. When ADAMTS-1 and fibulin-1 were transfected together, a reduction in the cell proliferation of both cell lines was observed. Specifically, in MDA-MB-231 cells, 35 and 48 Ki-67-positive nuclei were detected in the double transfectants and control cells, respectively, whereas in MCF-7 cells, 45 and 64 Ki-67-positive nuclei were detected in the double transfectants and control cells, respectively [Figure 4].
Figure 4. Coexpression of ADAMTS-1 and fibulin-1 reduces the proliferation of the breast cancer cell lines MCF-7 and MDA-MB-231. The proliferative state of breast cancer cell lines was measured by quantifying the proliferative nuclear marker Ki-67 by immunocytochemistry. Top Left: representative images of Ki-67 staining in the indicated MDA-MB-231 transfectants; bar, 10 μm. Top Right: quantification of Ki-67-positive nuclei (as a percentage of total nuclei labeled by DAPI staining). Bottom Left, representative images of Ki-67 staining in the indicated MCF-7 transfectants; bar, 10 μm. Bottom Right, quantification of Ki-67-positive nuclei (as a percentage of total nuclei labeled by DAPI staining)
Effects of the fibulin-1/ADAMTS-1 interaction on mammosphere formation
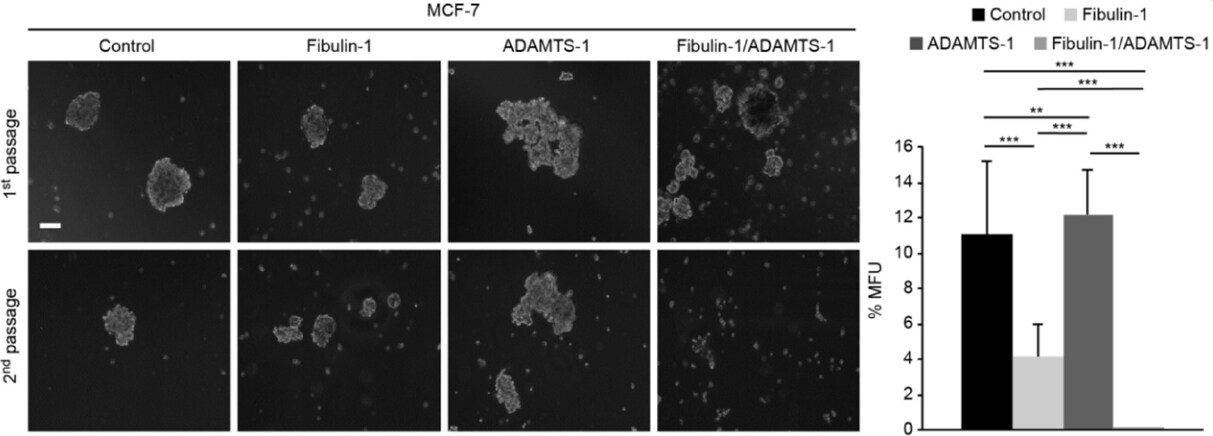
One way of mimicking a tissue environment using 3D cell culture in vitro is through the formation of mammospheres, allowing the quantification of the size of these structures as well as the self-renewal capacity of the cell. In this context, we were able to examine the influence of fibulin-1 and ADAMTS-1 on the mammosphere formation of breast cancer cells. The fibulin-1/ADAMTS-1 interaction resulted in fewer and smaller mammospheres than did the other three combinations [Figure 5]. Furthermore, significant differences in mammosphere forming units (MFUs) were observed among all possible combinations. It is remarkable that the ability to develop MFUs was almost completely abolished when fibulin-1 and ADAMTS-1 were coexpressed (no MFUs were detected). Compared with the control cells, cells expressing fibulin-1 also exhibited a diminished ability to develop MFUs (11 MFUs vs. 4 MFUs). In contrast, ADAMTS-1 expression slightly increased this capacity. Taken together, our data indicate that fibulin-1 and ADAMTS-1 affect the ability of this cell line to form mammospheres.
Figure 5. Coexpression of ADAMTS-1 and fibulin-1 abolishes mammosphere formation. Left: representative images of the mammospheres derived from MCF-7 transfectants in two consecutive generations. Control, MCF-7 cells transfected with an empty vector. Top row, mammosphere appearance before dissociation. Bottom row, mammosphere appearance 6 days after dissociation. Right: mammospheres were dissociated and passaged at a density of 20 cell/well in 96-well plates, and MFUs were counted and calculated as a percentage of the mammospheres formed from the number of cells seeded. White bar: 100 μm
Analysis of ADAMTS-1 expression in breast cancer tissue samples
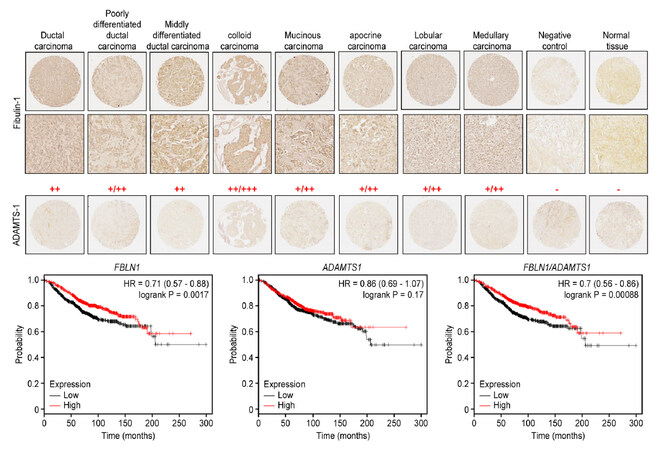
We performed an immunohistochemical analysis of the expression of ADAMTS-1 and fibulin-1 in a tissue array containing 69 breast cancer tissue samples, including 18 ductal carcinomas, 3 poorly differentiated ductal carcinomas, 12 moderately differentiated ductal carcinomas, 1 well-differentiated ductal carcinoma, 20 lobular carcinomas, 6 mucoid carcinomas, 3 colloid carcinomas, 3 apocrine carcinomas, and 3 medullary atypical carcinomas [Supplementary Table 1]. Overall, using serial slides, the results indicated no colocalization of the proteins regardless of the tumor type or stage [Figure 6]. Consistently, ADAMTS-1 immunoreactivity was almost absent or expressed at residual levels in all types of carcinoma samples [Figure 6]. On the other hand, positive immunostaining for fibulin-1 was distributed in both the tumor cells and the stromal spaces with the exception of the colloid carcinoma, in which fibulin-1 expression was restricted to the tumor cells [Figure 6, Top].
Figure 6. Different patterns of ADAMTS-1 and fibulin-1 expression in a breast cancer tissue array. Top: representative images of tissues used for the detection of ADAMTS-1 and fibulin-1 in different stages of breast carcinoma. Top two lines: fibulin-1 pattern at ×4 and ×20 magnifications. Bottom line: ADAMTS-1 pattern at ×4 magnification. Negative controls were performed by omitting primary antibodies. The intensity of fibulin-1 immunostaining is indicated as strong (+++), moderate (++), weak (+) or negative (-). Bottom: Kaplan-Meier overall survival plots for FBLN1, ADAMTS1 and FBLN1/ADAMTS1
We next assessed the association of ADAMTS1 and FBLN1 expression with clinical outcome using data available at www.kmplot.com. Overall survival Kaplan-Meier analysis revealed that the high expression of FBLN1 and ADAMTS1 correlated with the best prognosis for breast cancer patients (Figure 6, Bottom; measured as a mean of expression of both genes, P < 0.001). Similarly, high levels of FBLN1 can be considered a marker for good breast cancer prognosis (P < 0.005). On the other hand, this analysis also showed no influence of ADAMTS1 expression level on overall survival rate [Figure 6, Bottom]. We also evaluated progression-free survival and metastasis-free survival and found no significant differences in any of the cases analyzed (each gene alone or both together; data not shown). These data suggest that factors other than these two genes could modulate the pro- or anti-tumor activities of both fibulin-1 and ADAMTS-1.
Taken together, our results suggest that the inhibitory effect on the invasion, migration or mammosphere formation processes exhibited by breast cancer cell models cannot be fully extended to breast cancer tissue samples. Therefore, it cannot be ruled out that other factors can influence tumor cell behavior, such as the presence of protease inhibitors or the generation of bioactive fragments due to proteolytic activities. Additionally, the presence of fibulins other than fibulin-1, such as fibulin-5, can influence context-specific specific effects on breast cancer cells[37]. In this sense, fibulin-5 inhibits these same cellular properties; it was detected in a tissue array analysis of breast cancer samples and was inversely correlated with Ki-67 immunostaining[38].
Discussion
ECM proteins can be considered as important players in tissue homeostasis as it is considered the cellular component. Interactions among different components of the ECM have crucial roles in normal and pathological processes such as bone remodeling or tumor development[1,39]. The components of the ECM that contain multiple motifs within the same protein unit are good candidates to establish various interactions and build bridge connections between cells and other components of the ECM. Such is the case for the members of two extracellular families of proteins, the fibulins and ADAMTSs[4,21]. Members of both families have been associated with cancer development and progression and have been reported on many occasions to interact with other components of the ECM, acting as pro- or anti-tumor agents[18,23].
Fibulin-1 has been previously described to be an ADAMTS-1 cofactor and to stimulate the proteolytic activity of ADAMTS-1 towards aggrecan and versican[19,40]. Herein, we have been able to describe the significance of this fibulin-1/ADAMTS-1 interaction in the proliferation, migration and invasion properties of breast cancer cell lines. Overall, our data support the idea that ADAMTS-1 is a pro-tumor agent in breast cancer, whereas fibulin-1 is able to block this effect and acts as an anti-tumor agent. We have been able to show that the presence of both proteins reduces the migration and invasion properties of two breast cancer cell lines, MCF-7 and MDA-MB-231. The participation of ADAMTS-1 in cancer has been associated with mammary tumor growth and progression to metastasis in the MMTV-PyMT model in an Adamts1-/-background[10], whereas it has also been reported to be a tumor suppressor gene through proteolysis of nidogen-1 and nidogen-2 in an HEK293T tumor xenograft assay[41]. In our study, ADAMTS-1 expression was not detected by tissue array analysis in samples of different stages of breast cancer. Additionally, the Kaplan- Meier plots reflect the absence of an influence of ADAMTS-1 on expected patient overall survival. However, the ectopic expression of ADAMTS-1 in breast cancer cell lines increased the tumorigenic potential of these cell lines in terms of proliferation, migration, invasion and mammosphere formation. Initially, breast cancer profiles showed the downregulation of ADAMTS-1 in malignant tumors compared to normal tissues[42].
However, other studies indicated the participation of ADAMTS-1 in aiding breast cancer development[10,11,13,43]. Thus, the participation of ADAMTS-1 in breast cancer development and progression might depend on substrates it is able to act upon, such as versican to facilitate spreading of tumor cells or semaphorin 3C, which is known to promote metastatic potential of breast cancer cells[10,44]. Interestingly, ADAMTS1 was the third most overexpressed gene in highly metastatic MDA-MB-231 clones, and the knockdown of ADAMTS-1 in these clones regressed disease spread and reduced secondary tumor burden[45,46]. The effect of this protein might also be dependent on which cell type produces and secretes ADAMTS-1, similar to the paracrine effect of breast cancer cells over cancer-associated fibroblasts of the vicinity[15]. Finally, the impact of ADAMTS-1 might also depend on other ECM proteins that interact and bind with ADAMTS-1 and, as result, are able to modulate ADAMTS-1 activity.
The fact that fibulin-1 has been described as an ADAMTS-1 cofactor able to modulate ADAMTS-1 activity further implies the roles of both proteins in tumor development.
Fibulin-1 can interact with various ECM components, including known ADAMTS-1 substrates, such as nidogen-1 and versican[2,21,27]. Fibulin-1 has also been implicated in cellular transformation and, similar to ADAMTS-1, can behave as both a tumorigenic factor or tumor suppressor depending on the tissue environment[23]. Our results showed that the ectopic expression of fibulin-1 in breast cancer cell lines was able to block the tumorigenic properties of these cell lines. Furthermore, fibulin-1 expression was able to dramatically block these properties even in the presence of ADAMTS-1. Similar properties have also been described in our laboratory in the interaction of two related components of the ECM, fibulin-2 and ADAMTS-12. In this interaction, the expression of ADAMTS-12 alone exerted pro-tumoral activities, whereas the coexpression of both ADAMTS-12 and fibulin-2 showed an important anti-tumoral effect[47].
It is noteworthy that fibulin-1 expression was initially associated with improved survival in patients with a lymphoid infiltrate at the tumor site[48]. Similarly, the Kaplan-Meier analysis showed better prognosis for patients with high levels of fibulin-1 than for those with low fibulin-1 expression. In fact, the low expression of fibulin-1 has been described in various tumor types, such as gastric cancer, bladder cancer, colorectal cancer and hepatocellular carcinoma. In all these cases, low expression correlates with the hypermethylation of the FBLN1 promoter and can be used as a prognosis marker[30-33,49]. Thus, our data are in accordance with all these studies in which it seems feasible that the presence of fibulin-1 might have a prevalent antitumor activity. In our immunohistochemical analysis, we could detect the presence of only fibulin-1; no ADAMTS-1 was observed. From our cellular experiments, it is clear that the presence of both proteins has a pronounced effect on the tumorigenic properties of both cell lines. The downregulation of ADAMTS-1 might be a mechanism of escape towards tumor progression, avoiding binding with fibulin-1 and thus eliminating the anti-tumoral properties of their interaction.
In summary, the interactions between members of both families of ECM proteins, fibulins and ADAMTSs, seem to be important for cancer development, as is the case for ADAMTS-12/fibulin-2 and ADAMTS-1/fibulin-1. Furthermore, we have also been able to demonstrate the cleavage of fibulin-2 by ADAMTS-4 and ADAMTS-5, which increased the tumorigenic potential of breast cancer cell lines[50]. Therefore, it is tempting to speculate the existence of similar mechanisms that control fibulin-1 participation in breast cancer development and its interaction with other ECM components, a hypothesis that warrant further study.
Declarations
AcknowledgmentsWe also thank the IUOPA units of “Banco de tumores” and “ Histopatología molecular en modelos animales en cancer”.
Authors’ contributionsMade substantial contributions to the conception and design of the study and performed data analysis and interpretation: Mohamedi Y, Fontanil T, Cal S, Obaya AJ
Performed data acquisition: Cobo T, Cobo JL
Provided technical support: García-Suárez O
Provided administrative support and performed histological interpretation: Vega JA, Cobo J
Wrote the manuscript: Mohamedi Y, Cal S, Obaya AJ
Availability of data and materialsNot applicable.
Financial support and sponsorshipThis work was supported by the Instituto Asturiano de Odontología (IAO). Mohamedi Y is recipient of a fellowship from the FICYT, “Fundación para el Fomento en Asturias de la Investigación Científica Aplicada y la Tecnología” (“Severo Ochoa” Research Program, Principado de Asturias), and Tania Fontanil is recipient of a contract from the “Departamento de Investigación de Clínica Órdoñez (Oviedo)”.
Conflicts of interestAll authors declared that there are not conflicts of interest.
Ethical approval and consent to participateThe study methodologies conformed to the standards set by the Declaration of Helsinki and were approved by the Ethics Committee of Clinical Investigation of the Hospital Universitario de Asturias (HUCA).
Consent for publicationNot applicable.
Copyright© The Author(s) 2019.
REFERENCES
1. Binder MJ, McCoombe S, Williams ED, McCulloch DR, Ward AC. The extracellular matrix in cancer progression: role of hyalectan proteoglycans and ADAMTS enzymes. Cancer Lett 2017;385:55-64.
2. Tsuda T. Extracellular interactions between fibulins and transforming growth factor (TGF)-beta in physiological and pathological conditions. Int J Mol Sci 2018;19:E2787.
3. Spaw M, Anant S, Thomas SM. Stromal contributions to the carcinogenic process. Mol Carcinog 2017;56:1199-213.
4. Kelwick R, Desanlis I, Wheeler GN, Edwards DR. The ADAMTS (a disintegrin and metalloproteinase with thrombospondin motifs) family. Genome Biol 2015;16:113.
5. Kuno K, Kanada N, Nakashima E, Fujiki F, Ichimura F, et al. Molecular cloning of a gene encoding a new type of metalloproteinase-disintegrin family protein with thrombospondin motifs as an inflammation associated gene. J Biol Chem 1997;272:556-62.
6. Shindo T, Kurihara H, Kuno K, Yokoyama H, Wada T, et al. ADAMTS-1: a metalloproteinase-disintegrin essential for normal growth, fertility, and organ morphology and function. J Clin Invest 2000;105:1345-52.
7. Iruela-Arispe ML, Carpizo D, Luque A. ADAMTS1: a matrix metalloprotease with angioinhibitory properties. Ann N Y Acad Sci 2003;995:183-90.
8. Brown HM, Dunning KR, Robker RL, Boerboom D, Pritchard M, et al. ADAMTS1 cleavage of versican mediates essential structural remodeling of the ovarian follicle and cumulus-oocyte matrix during ovulation in mice. Biol Reprod 2010;83:549-57.
9. Tan Ide A, Ricciardelli C, Russell DL. The metalloproteinase ADAMTS1: a comprehensive review of its role in tumorigenic and metastatic pathways. Int J Cancer 2013;133:2263-76.
10. Ricciardelli C, Frewin KM, tan Ide A, Williams ED, Opeskin K, et al. The ADAMTS1 protease gene is required for mammary tumor growth and metastasis. Am J Pathol 2011;179:3075-85.
11. Liu YJ, Xu Y, Yu Q. Full-length ADAMTS-1 and the ADAMTS-1 fragments display pro- and antimetastatic activity, respectively. Oncogene 2006;25:2452-67.
12. Obika M, Ogawa H, Takahashi K, Li J, Hatipoglu OF, et al. Tumor growth inhibitory effect of ADAMTS1 is accompanied by the inhibition of tumor angiogenesis. Cancer Sci 2012;103:1889-97.
13. Hirano T, Hirose K, Sakurai K, Makishima M, Sasaki K, et al. Inhibition of tumor growth by antibody to ADAMTS1 in mouse xenografts of breast cancer. Anticancer Res 2011;31:3839-42.
14. Freitas VM, do Amaral JB, Silva TA, Santos ES, Mangone FR, et al. Decreased expression of ADAMTS-1 in human breast tumors stimulates migration and invasion. Mol Cancer 2013;12:2.
15. Tyan SW, Hsu CH, Peng KL, Chen CC, Kuo WH, et al. Breast cancer cells induce stromal fibroblasts to secrete ADAMTS1 for cancer invasion through an epigenetic change. PLoS One 2012;7:e35128.
16. Ham SA, Yoo T, Lee WJ, Hwang JS, Hur J, et al. ADAMTS1-mediated targeting of TSP-1 by PPARdelta suppresses migration and invasion of breast cancer cells. Oncotarget 2017;8:94091-103.
17. Li M, Liu L, Zang W, Wang Y, Du Y, et al. miR365 overexpression promotes cell proliferation and invasion by targeting ADAMTS-1 in breast cancer. Int J Oncol 2015;47:296-302.
19. Lee NV, Rodriguez-Manzaneque JC, Thai SN, Twal WO, Luque A, et al. Fibulin-1 acts as a cofactor for the matrix metalloprotease ADAMTS-1. J Biol Chem 2005;280:34796-804.
20. Argraves WS, Dickerson K, Burgess WH, Ruoslahti E. Fibulin, a novel protein that interacts with the fibronectin receptor beta subunit cytoplasmic domain. Cell 1989;58:623-9.
21. de Vega S, Iwamoto T, Yamada Y. Fibulins: multiple roles in matrix structures and tissue functions. Cell Mol Life Sci 2009;66:1890-902.
23. Obaya AJ, Rua S, Moncada-Pazos A, Cal S. The dual role of fibulins in tumorigenesis. Cancer Lett 2012;325:132-8.
24. Greene LM, Twal WO, Duffy MJ, McDermott EW, Hill AD, et al. Elevated expression and altered processing of fibulin-1 protein in human breast cancer. Br J Cancer 2003;88:871-8.
25. Twal WO, Czirok A, Hegedus B, Knaak C, Chintalapudi MR, et al. Fibulin-1 suppression of fibronectin-regulated cell adhesion and motility. J Cell Sci 2001;114:4587-98.
26. Qing J, Maher VM, Tran H, Argraves WS, Dunstan RW, et al. Suppression of anchorage-independent growth and matrigel invasion and delayed tumor formation by elevated expression of fibulin-1D in human fibrosarcoma-derived cell lines. Oncogene 1997;15:2159-68.
27. Timpl R, Sasaki T, Kostka G, Chu ML. Fibulins: a versatile family of extracellular matrix proteins. Nat Rev Mol Cell Biol 2003;4:479-89.
28. Moll F, Katsaros D, Lazennec G, Hellio N, Roger P, et al. Estrogen induction and overexpression of fibulin-1C mRNA in ovarian cancer cells. Oncogene 2002;21:1097-107.
29. Sadlonova A, Bowe DB, Novak Z, Mukherjee S, Duncan VE, et al. Identification of molecular distinctions between normal breast-associated fibroblasts and breast cancer-associated fibroblasts. Cancer Microenviron 2009;2:9-21.
30. Xu Z, Chen H, Liu D, Huo J. Fibulin-1 is downregulated through promoter hypermethylation in colorectal cancer: a CONSORT study. Medicine (Baltimore) 2015;94:e663.
31. Cheng YY, Jin H, Liu X, Siu JM, Wong YP, et al. Fibulin 1 is downregulated through promoter hypermethylation in gastric cancer. Br J Cancer 2008;99:2083-7.
32. Kanda M, Nomoto S, Okamura Y, Hayashi M, Hishida M, et al. Promoter hypermethylation of fibulin 1 gene is associated with tumor progression in hepatocellular carcinoma. Mol Carcinog 2011;50:571-9.
33. Xiao W, Wang J, Li H, Xia D, Yu G, et al. Fibulin-1 is epigenetically down-regulated and related with bladder cancer recurrence. BMC Cancer 2014;14:677.
34. Fontanil T, Mohamedi Y, Cal S, Obaya AJ. Assessing the influence of a protease in cell migration using the barrier-migration assay. Methods Mol Biol 2018;1731:133-43.
35. Dontu G, Abdallah WM, Foley JM, Jackson KW, Clarke MF, et al. In vitro propagation and transcriptional profiling of human mammary stem/progenitor cells. Genes Dev 2003;17:1253-70.
36. Gyorffy B, Benke Z, Lanczky A, Balazs B, Szallasi Z, et al. RecurrenceOnline: an online analysis tool to determine breast cancer recurrence and hormone receptor status using microarray data. Breast Cancer Res Treat 2012;132:1025-34.
37. Schiemann WP, Blobe GC, Kalume DE, Pandey A, Lodish HF. Context-specific effects of fibulin-5 (DANCE/EVEC) on cell proliferation, motility, and invasion. Fibulin-5 is induced by transforming growth factor-beta and affects protein kinase cascades. J Biol Chem 2002;277:27367-77.
38. Mohamedi Y, Fontanil T, Solares L, Garcia-Suarez O, Garcia-Piqueras J, et al. Fibulin-5 downregulates Ki-67 and inhibits proliferation and invasion of breast cancer cells. Int J Oncol 2016;48:1447-56.
39. Alford AI, Kozloff KM, Hankenson KD. Extracellular matrix networks in bone remodeling. Int J Biochem Cell Biol 2015;65:20-31.
40. Kern CB, Twal WO, Mjaatvedt CH, Fairey SE, Toole BP, et al. Proteolytic cleavage of versican during cardiac cushion morphogenesis. Dev Dyn 2006;235:2238-47.
41. Martino-Echarri E, Fernandez-Rodriguez R, Rodriguez-Baena FJ, Barrientos-Duran A, Torres-Collado AX, et al. Contribution of ADAMTS1 as a tumor suppressor gene in human breast carcinoma. Linking its tumor inhibitory properties to its proteolytic activity on nidogen-1 and nidogen-2. Int J Cancer 2013;133:2315-24.
42. Porter S, Scott SD, Sassoon EM, Williams MR, Jones JL, et al. Dysregulated expression of adamalysin-thrombospondin genes in human breast carcinoma. Clin Cancer Res 2004;10:2429-40.
43. Bonuccelli G, Casimiro MC, Sotgia F, Wang C, Liu M, et al. Caveolin-1 (P132L), a common breast cancer mutation, confers mammary cell invasiveness and defines a novel stem cell/metastasis-associated gene signature. Am J Pathol 2009;174:1650-62.
44. Esselens C, Malapeira J, Colome N, Casal C, Rodriguez-Manzaneque JC, et al. The cleavage of semaphorin 3C induced by ADAMTS1 promotes cell migration. J Biol Chem 2010;285:2463-73.
45. Kang Y, Siegel PM, Shu W, Drobnjak M, Kakonen SM, et al. A multigenic program mediating breast cancer metastasis to bone. Cancer Cell 2003;3:537-49.
46. Lu X, Wang Q, Hu G, van Poznak C, Fleisher M, et al. ADAMTS1 and MMP1 proteolytically engage EGF-like ligands in an osteolytic signaling cascade for bone metastasis. Genes Dev 2009;23:1882-94.
47. Fontanil T, Rua S, Llamazares M, Moncada-Pazos A, Quiros PM, et al. Interaction between the ADAMTS-12 metalloprotease and fibulin-2 induces tumor-suppressive effects in breast cancer cells. Oncotarget 2014;5:1253-64.
48. Pupa SM, Argraves WS, Forti S, Casalini P, Berno V, et al. Immunological and pathobiological roles of fibulin-1 in breast cancer. Oncogene 2004;23:2153-60.
49. Feng L, Yao C, Li P, Feng Y, Wang F, et al. Low expression of fibulin-1 correlates with unfavorable prognosis in gastric cancer. Tumour Biol 2016;37:9399-410.
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Mohamedi Y, Fontanil T, Cobo T, Vega JA, Cobo JL, García-Suárez O, Cobo J, Cal S, Obaya J. The molecular interaction of ADAMTS-1 and fibulin-1 and its potential contribution to breast cancer biology. J Cancer Metastasis Treat 2019;5:37. http://dx.doi.org/10.20517/2394-4722.2018.81
AMA Style
Mohamedi Y, Fontanil T, Cobo T, Vega JA, Cobo JL, García-Suárez O, Cobo J, Cal S, Obaya J. The molecular interaction of ADAMTS-1 and fibulin-1 and its potential contribution to breast cancer biology. Journal of Cancer Metastasis and Treatment. 2019; 5: 37. http://dx.doi.org/10.20517/2394-4722.2018.81
Chicago/Turabian Style
Mohamedi, Yamina, Tania Fontanil, Teresa Cobo, José A. Vega, Juan L. Cobo, Olivia García-Suárez, Juan Cobo, Santiago Cal, Álvaro J. Obaya. 2019. "The molecular interaction of ADAMTS-1 and fibulin-1 and its potential contribution to breast cancer biology" Journal of Cancer Metastasis and Treatment. 5: 37. http://dx.doi.org/10.20517/2394-4722.2018.81
ACS Style
Mohamedi, Y.; Fontanil T.; Cobo T.; Vega JA.; Cobo JL.; García-Suárez O.; Cobo J.; Cal S.; Obaya J. The molecular interaction of ADAMTS-1 and fibulin-1 and its potential contribution to breast cancer biology. J. Cancer. Metastasis. Treat. 2019, 5, 37. http://dx.doi.org/10.20517/2394-4722.2018.81
About This Article
Special Issue
Copyright
Data & Comments
Data
 Cite This Article 24 clicks
Cite This Article 24 clicks












Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.