Association between the cytotoxic T-lymphocyte antigen-4 polymorphisms and breast cancer risk and prognosis
Abstract
Aim: The aim was to evaluate the potential influences of cytotoxic T-lymphocyte antigen-4 (CTLA-4) gene polymorphisms on breast cancer risk, the distribution of CTLA-4 single nucleotide polymorphisms (1661AG) in breast cancer patients and control subjects was investigated.
Methods: In this case-control study, 100 patients with breast cancer as case group and 100 healthy participants as a control group were compared. Genotypes were determined by the polymerase chain reaction-restriction fragment length polymorphism method. Demographic characteristics of the study population, as well as tumor size, tumor grade and stage were collected in a questionnaire designed for this study. The collected data were statistically analyzed by SPSS-16.0 (SPSS Inc., Chicago, USA) predictive analytic software using the Chi-square test.
Results: The mean age of women was 43.42 ± 13.1 years. The AA genotype was frequent in case group (43%) whereas the AG genotype was found more in the control group (69%). There was no significant relationship between the studied polymorphisms and the grade, stage and size of the tumor, nor between the studied polymorphisms and estrogen receptor, progesterone receptor and lymph node involvement (P > 0.05). Significant association between the studied polymorphisms and breast cancer metastases was found (P = 0.02).
Conclusion: According to the results of the study, the AA genotype is associated with breast cancer, but none of the studied gene polymorphisms is associated with prognostic factors such as tumor stage, grade or size.
Keywords
Introduction
Breast cancer is the most common cancer in women (30% of all cancers among women in developed countries), but is treatable if early diagnosis and treatment occur.[1] The incidence of breast cancer has constantly increased since 1940[2] and based on WHO reports, there is a 2% annual increase in breast cancer prevalence.[3] There are no exact statistics on the prevalence of breast cancer among the Iranian population (the source of our study); estimates show that Iran has moderate, but increasing, prevalence.[4] Both genetic and environmental susceptibilities are included in breast cancer etiology,[5-8] but the exact etiology has not been definitively identified. Current studies confirm the role of the immune system on the etiology of breast cancer.
It was shown that cancer cells provoke immune recognition, but the biologic importance of antitumor innate and adaptive responses, which are frequently detected in cancer-bearing hosts, remains incompletely understood.[9] The most considerable antitumor response is made by human cellular immunity mediated by T-lymphocyte and natural killer (NK) cells. It follows that variants of genes included in the regulation, and proliferation of T-lymphocyte and NK cells would be effective in predicting the risk of breast cancer. Cytotoxic T-lymphocyte antigen-4 (CTLA-4) is coded by a gene on chromosome 2q33. It is a member of the immunoglobulin super family, which transmits an inhibitory signal to T cells. CTLA-4 binds to B-7 on antigen-presenting cells, and polymorphism of CTLA-4 gene interferes with surface activity of B-7, preventing T-lymphocyte from activating.[10,11] In fact, CTLA-4 prevents immune response[12] and its tumor-killing activity.[13]CTLA-4 gene is composed of 4 exons and possibly plays a signifi cant role in diseases related to T cells. More than 100 single-nucleotide polymorphisms are recognized on the CTLA-4 gene. Among them, AG dysmorphisms, located on +49 of exon 1, could make amino acid (threonine into alanine) on CTLA-4 protein.[14]
Current studies show that this polymorphism affects the ability of CTLA-4 to bind to B-7 cells and to activate T cells.[15,16] These surveys show that translocation of A allele to G allele on +49 zone decreases the role of CTLA-4 on T-cell responses,[15-18] although there are contradictory reports on the relationship between +49 A to G polymorphisms and cancer development.
Higher expression of CTLA-4 is seen in persons with thiamine on zone -318 of CTLA-4 gene promoter or homogenous adenine on exon 1 of codon +49.[14] There have been several studies on the relationship between polymorphisms on CTLA-4 gene and autoimmune diseases such as graves, diabetes mellitus type one, lupus and Hashimoto thyroiditis[19-22] and the tendency to develop cancer.[23-27] Results of some studies show an inverse relation between polymorphisms of autoimmune diseases and malignancies on CTLA-4 gene. Alleles discovered in autoimmune diseases are not seen in malignancies or are related to a good prognosis of cancers. In a study in Iran, results suggested higher risk of breast cancer among AA and AG genotypes on +49 zone, but there was no difference between -318 CT and -1666 AG among case and control groups.[23,24]
Considering the high prevalence of breast cancer and also some confi rmed evidence of a relationship between CTLA-4 gene polymorphisms and breast cancer, we conducted this study to assess the relationship between CTLA-4 gene polymorphisms and both incidence and clinic pathologic features of breast cancer. The results of this study would help physicians to recognize the prognosis and risk ratio of patients with a high risk of breast cancer.
Methods
Study subjects
The study group consisted of a total of 100 Iranian women with breast cancer and 100 healthy cancer-free control individuals. Informed consent was obtained from each subject, and each participant was then interviewed to collect detailed information on demographic characteristics such as sex and age. Some clinic pathologic features of breast cancer patients, such as tumor size, lymph node involvement, tumor type, and estrogen receptor (ER), were also obtained from their medical files [Table 1].
Relation between CTLA-4 gene polymorphisms and tumor grade, stage, size, metastasis, estrogen receptor, progesterone receptor and age
| AA (%) | AG (%) | GG (%) | P | |
|---|---|---|---|---|
| Tumor grade | ||||
| 1 | 20 (80) | 23 (71.9) | 1 (33.3) | 0.21 |
| 2 | 3 (12) | 5 (15.6) | 1 (33.3) | |
| 3 | 2 (8) | 4 (12.5) | 1 (33.3) | |
| Tumor stage | ||||
| ≤ 2 | 15 (75) | 18 (72) | 3 (100) | 0.50 |
| > 2 | 5 (25) | 7 (28) | 0 (0) | |
| Tumor size | ||||
| ≤ 2 cm | 11 (68.8) | 14 (58.3) | 0 (0) | 0.50 |
| 2-5 cm | 3 (18.8) | 8 (33.3) | 1 (100) | |
| > 5 cm | 2 (12.5) | 2 (8.3) | 0 (0) | |
| Metastasis | ||||
| No | 27 (64.3) | 43 (87.7) | 5 (83.3) | 0.02 |
| Yes | 15 (35.7) | 6 (12.2) | 1 (16.7) | |
| Estrogen receptor | ||||
| Negative | 10 (37) | 9 (23.7) | 2 (33.3) | 0.49 |
| Positive | 17 (63) | 29 (76.3) | 4 (66.7) | |
| Progesterone receptor | ||||
| Negative | 12 (44.4) | 8 (21.6) | 2 (33.3) | 0.15 |
| Positive | 15 (55.6) | 29 (78.4) | 4 (66.7) | |
| Lymph node involvement | ||||
| No | 36 (83.7) | 38 (74.5) | 5 (83.3) | 0.53 |
| Yes | 7 (16.3) | 13 (25.5) | 1 (16.7) | |
| Age | ||||
| Under 40 years | 24 (29.3) | 54 (65.9) | 4 (4.9) | 0.30 |
| Over 40 years | 47 (29.8) | 66 (55.9) | 5 (4.2) | |
Patients were recruited between February 2013 and October 2014 at the Shahid Sadughi Hospital and Cancer Hospital, Yazd, Iran. Control subjects were cancer-free individuals, and they were randomly selected from the same regions and the same time period as the patients were collected. The selection criteria included no individual history of breast or other cancers.
This study was approved by the Ethics Committee of Shahid Sadoughi University of Medical Sciences, Yazd, Iran. A written informed consent was taken from all patients.
Polymorphism genotyping
Peripheral blood (5 mL) was collected from subjects after informed consent was obtained. Genomic DNA was extracted from peripheral blood using the DNA extraction kit (BioFlux, cat: BSC 06M1, Hangzhou, Bioer Technology Co., Ltd, China).
Genotyping was performed by polymerase chain reaction (PCR)-restriction fragment length polymorphism method. The polymorphic region was amplifi ed by PCR using the following primers: (forward) 5’-CTAAGAGCATCCGCTTGCACCT-3’ and (reverse) 5’-TTGGTGTGATGCACAGAAGCCTTT-3’ in a 25 μL reaction solution containing 0.3 μg of genomic DNA, ×1 PCR buffer, 0.3 mmol/L MgCl2, 0.2 mmol/L dNTPs, 2 U tag DNA polymerase and 0.1 μmol/L of each primer.
The following PCR program was run: 94 °C for 4 min, 30 cycles of 94 °C for 30 s, 58 °C for 30 s and 7 °C for 45 s. Final extension was carried out at 72 °C for 5 min. The lengths of the PCR products were 486 bp (1661AG).
The PCR products were digested with restriction enzymes Tru1I (MseI) according to the manufacturer’s instructions (Thermo Scientific Fermentas, USA) and analyzed by 2% agarose gel electrophoresis. The cut site for Tru1I (MseI) was 5’-TTAA-3’.
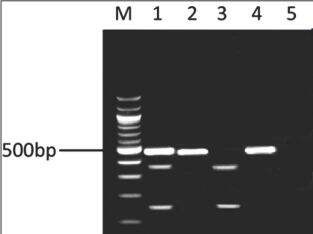
The digested fragments in 1661AG were 139 and 347 bp. Presence of the A allele was recognized by detecting digested 347 and 139 bp fragments on gel, and the G allele by detecting intact primary 486 bp band [Figure 1]. Comparisons of genotype and allele frequencies in cases and controls were assessed by Chi-square and t-test using SPSS-16.0 (SPSS Inc., Chicago, USA). Statistical software and statistical significance were set at P ≤ 0.05. The odds ratio and 95% confidence interval were also calculated.
Results
A total of 100 women with breast cancer and 100 healthy controls were enrolled in this study. Mean age of the case group was 48.92 ± 9.85 years and of the control group was 37.92 ± 13.67 years (P < 0.001; t-test). This study was done in patients who presented at Shahid Sadughi Hospital and Cancer Hospital, Yazd, Iran. The grade, stage and size of the tumor in the case group are shown in Table 1. About 73.3% of patients had grade one tumors, and about 75% had stages 1 and 2 tumors. Tumor size in more than 60.9% of patients was ≤ 2 mm.
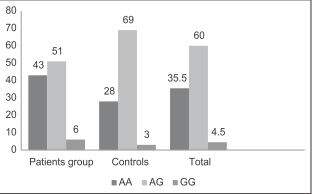
Polymorphisms of CTLA-4 gene are shown in [Figure 2]. There is a significant relationship between groups according to gene polymorphisms (P = 0.03). Frequency of AA polymorphisms in the case group is higher than in controls whereas AG polymorphisms are more frequent in the control group.
Figure 2. Frequency of AA, AG and GG polymorphisms of cytotoxic T-lymphocyte antigen-4 gene in patients, controls and total study group, showing that AA genotype in patients is more than this genotype in controls while AG polymorphism is more frequent in the control group. The Y-axis is showing the number of the cases and the X-axis is showing different types of genotypes
Table 1 shows polymorphisms according to tumor stage. There is no significant difference among study groups according to tumor grade (P = 0.21). Also there was no relationship between polymorphism and tumor stage (P = 0.50) and tumor size (P = 0.50).
Considering the difference in mean age between the two study groups, we analyzed polymorphisms of CTLA-4 gene in all participants according to age groups of under or over 40 years, but found no significant difference (P = 0.30) [Table 1]. We determined that the age difference between the two groups had no confounding effect on the study results.
Analysis of metastasis showed that there is a significant relationship between CTLA-4 gene polymorphism and metastasis. Patients with AA genotype had higher rates of metastasis (P = 0.02) [Table 1].
Analysis also showed no relationship between CTLA-4 gene polymorphisms and ER (P = 0.49), progesterone receptor (P = 0.15) and lymph node involvement (P = 0.53) [Table 1].
Discussion
There is increasing attention to the relationship between several genes’ polymorphisms and polygenetic diseases such as hypertension, diabetes, and various malignancies.[28] T cells and NK cells have a substantial role in working against tumors.[29] T-lymphocyte, especially T killer cell, is the most important in defending cells against tumors. CTLA-4 molecule expresses on T-lymphocyte as an inhibitor and plays different roles in T-cell activity. It could inhibit amplification of T cells or even induce apoptosis of activated T cells.[30]CTLA-4-mediated suppression of tumor immunity has been previously reported.[13] Several studies have demonstrated the effect of CTLA-4 blockade in enhancing immunity to tumors.[14-16] There are some studies on the relationship between CTLA-4 gene polymorphisms and breast cancer, but results were contradictory.[31] In order to clarify the role of genetic variants of CTLA-4 gene in immune suppression of patients with cancer, the distribution of CTLA-4 gene single nucleotide polymorphisms (1661AG), in breast cancer patients and control subjects, were investigated and their associations were assessed with prognostic factors. The present case control study was done to determine possible relationships between AA, AG and GG polymorphisms of CTLA-4 gene and breast cancer-related factors. Results revealed that the frequency of AA and GG genotypes in breast cancer patients is higher than in controls, while AG genotype is more frequent in healthy controls. These results confirm the findings of two other studies in Iran[23] and China,[32] which found GG genotype is more frequent in breast cancer patients. However, Erfani et al.[24] did not find any difference between their study groups in terms of AG genotype. Another study in China suggested that AG genotype is more prevalent in breast cancer patients.[33] Furthermore, Sun et al.[15] reported that T cells with AA genotype are less active than those with GG genotype, and AG is related to different cancer incidence in humans.
The results of our study did not find any relation between AA, AG and GG genotypes with respect to tumor stage, grade, receptors or lymph node involvement. These results are in agreement with results of Wang et al.,[16] who found no significant relationship between AG genotype and tumor size and lymph node involvement. Erfani et al.[24] found a relationship between AA genotype and lower lymph node involvement and higher ER expression, but Ghaderi et al.[23] found that AA genotype is related to higher rates of lymph node involvement and tumor size; these findings are different from our findings. Bi et al.[34] in his study reported that CTLA-4 expression is higher in stage 2 than stage 3 patients. There are also some other studies, which revealed that CTLA-4 gene polymorphisms are related to higher stages and lymph node metastasis,[6,32] which is not consistent with our study. According to age, in our study and also in the Bi et al.,[34] there was no relationship between CTLA-4 gene polymorphisms and age.
Li et al.[32] found a relationship between all CTLA-4 gene polymorphisms with estrogen and progesterone receptors. Also, Erfani et al.[24] detected a relationship between AG genotype and ER expression, which was not consistent with our findings.
One of the limitations of our study is the failure to take into account risk factors such as age at menarche, menopausal status, and environmental factors. It is important to investigate the interaction between single-nucleotide polymorphisms and these factors on the risk of breast cancer in a larger sample size in further studies.
Based on our study, there is a relationship between CTLA-4 gene 1661AG polymorphisms and incidence of breast cancer, but these polymorphisms are not effective for prognosis. Considering the controversial reports on this issue, more studies are needed with larger sample size. Also, a critical review and possible meta-analysis of present studies are needed to make an exact estimation of the results of current studies.
Financial support and sponsorship
Nil.
Conflicts of interest
There are no conflicts of interest.
REFERENCES
1. Boring CC, Squires TS, Tong T, Montgomery S. Cancer statistic, 2004. CA Cancer J Clin 1994;44:7-26.
2. Lippman ME, Lichter AS, Danforth DN Jr. Diagnosis and Management of Breast Cancer. Philadelphia: WB Saunders Co.; 1988.
3. Henderson IC, Canellos GP. Cancer of the breast: the past decade (first of two parts). N Engl J Med 1980;302:17-30.
4. Bakhtiari A, Haj-Ahmadi M. Five-year assessment of breast cancer at Rajaee Hospital, Baboolsar (1991-1996). Iran J Obstets Gynecol Infertil 2006;9:47-52.
5. Li F, Sturgis EM, Chen X, Zafereo ME, Wei Q, Li G. Association of p53 codon 72 polymorphism with risk of second primary malignancy in patients with squamous cell carcinoma of the head and neck. Cancer 2010;116:2350-9.
6. Wang G, Yu D, Tan W, Zhao D, Wu C, Lin D. Genetic polymorphism in chemokine CCL22 and susceptibility to Helicobacter pylori infection-related gastric carcinoma. Cancer 2009;115:2430-7.
7. Kristensen VN, Børresen-Dale AL. SNPs associated with molecular subtypes of breast cancer: On the usefulness of stratified Genome-wide Association Studies (GWAS) in the identification of novel susceptibility loci. Mol Oncol 2008;2:12-5.
8. Kraft P, Haiman CA. GWAS identifies a common breast cancer risk allele among BRCA1 carriers. Nat Genet 2010;42:819-20.
10. Hurwitz AA, Kwon ED, van Elsas A. Costimulatory wars: the tumor menace. Curr Opin Immunol 2000;12:589-96.
11. Chen L. Co-inhibitory molecules of the B7-CD28 family in the control of T-cell immunity. Nat Rev Immunol 2004;4:336-47.
12. Leach DR, Krummel MF, Allison JP. Enhancement of antitumor immunity by CTLA-4 blockade. Science 1996;271:1734-6.
13. Hurwitz AA, Foster BA, Kwon ED, Truong T, Choi EM, Greenberg NM, Burg MB, Allison JP. Combination immunotherapy of primary prostate cancer in a transgenic mouse model using CTLA-4 blockade. Cancer Res 2000;60:2444-8.
14. Ligers A, Teleshova N, Masterman T, Huang WX, Hillert J. CTLA-4 gene expression is influenced by promoter and exon 1 polymorphisms. Genes Immun 2001;2:145-52.
15. Sun T, Zhou Y, Yang M, Hu Z, Tan W, Han X, Shi Y, Yao J, Guo Y, Yu D, Tian T, Zhou X, Shen H, Lin D. Functional genetic variations in cytotoxic T-lymphocyte antigen 4 and susceptibility to multiple types of cancer. Cancer Res 2008;68:7025-34.
16. Wang L, Li D, Fu Z, Li H, Jiang W, Li D. Association of CTLA-4 gene polymorphisms with sporadic breast cancer in Chinese Han population. BMC Cancer 2007;7:173.
17. Kouki T, Sawai Y, Gardine CA, Fisfalen ME, Alegre ML, DeGroot LJ. CTLA-4 gene polymorphism at position 49 in exon 1 reduces the inhibitory function of CTLA-4 and contributes to the pathogenesis of Graves' disease. J Immunol 2000;165:6606-11.
18. Mäurer M, Loserth S, Kolb-Mäurer A, Ponath A, Wiese S, Kruse N, Rieckmann P. A polymorphism in the human cytotoxic T-lymphocyte antigen 4 (CTLA4) gene (exon 1 +49) alters T-cell activation. Immunogenetics 2002;54:1-8.
19. Ueda H, Howson JM, Esposito L, Heward J, Snook H, Chamberlain G, Rainbow DB, Hunter KM, Smith AN, Di Genova G, Herr MH, Dahlman I, Payne F, Smyth D, Lowe C, Twells RC, Howlett S, Healy B, Nutland S, Rance HE, Everett V, Smink LJ, Lam AC, Cordell HJ, Walker NM, Bordin C, Hulme J, Motzo C, Cucca F, Hess JF, Metzker ML, Rogers J, Gregory S, Allahabadia A, Nithiyananthan R, Tuomilehto-Wolf E, Tuomilehto J, Bingley P, Gillespie KM, Undlien DE, Rønningen KS, Guja C, Ionescu-Tîrgovişte C, Savage DA, Maxwell AP, Carson DJ, Patterson CC, Franklyn JA, Clayton DG, Peterson LB, Wicker LS, Todd JA, Gough SC. Association of the T-cell regulatory gene CTLA4 with susceptibility to autoimmune disease. Nature 2003;423:506-11.
20. Han SZ, Zhang SH, Li R, Zhang WY, Li Y. The common -318C/T polymorphism in the promoter region of CTLA4 gene is associated with reduced risk of ophthalmopathy in Chinese Graves' patients. Int J Immunogenet 2006;33:281-7.
21. Anjos SM, Polychronakos C. Functional evaluation of the autoimmunity-associated CTLA4 gene: The effect of the (AT) repeat in the 3'untranslated region (UTR). J Autoimmun 2006;27:105-9.
22. Barreto M, Santos E, Ferreira R, Fesel C, Fontes MF, Pereira C, Martins B, Andreia R, Viana JF, Crespo F, Vasconcelos C, Ferreira C, Vicente AM. Evidence for CTLA4 as a susceptibility gene for systemic lupus erythematosus. Eur J Hum Genet 2004;12:620-6.
23. Ghaderi A, Yeganeh F, Kalantari T, Talei AR, Pezeshki AM, Doroudchi M, Dehaghani AS. Cytotoxic T lymphocyte antigen-4 gene in breast cancer. Breast Cancer Res Treat 2004;86:1-7.
24. Erfani N, Razmkhah M, Talei AR, Pezeshki AM, Doroudchi M, Monabati A, Ghaderi A. Cytotoxic T lymphocyte antigen-4 promoter variants in breast cancer. Cancer Genet Cytogenet 2006;165:114-20.
25. Zheng C, Huang D, Liu L, Björkholm M, Holm G, Yi Q, Sundblad A. Cytotoxic T-lymphocyte antigen-4 microsatellite polymorphism is associated with multiple myeloma. Br J Haematol 2001;112:216-8.
26. Monne M, Piras G, Palmas A, Arru L, Murineddu M, Latte G, Noli A, Gabbas A. Cytotoxic T-lymphocyte antigen-4 (CTLA-4) gene polymorphism and susceptibility to non-Hodgkin's lymphoma. Am J Hematol 2004;76:14-8.
27. Mao H, Zhang L, Yang Y, Zuo W, Bi Y, Gao W, Deng B, Sun J, Shao Q, Qu X. New insights of CTLA-4 into its biological function in breast cancer. Curr Cancer Drug Targets 2010;10:728-36.
29. Linsley PS, Ledbetter JA. The role of the CD28 receptor during T cell responses to antigen. Annu Rev Immunol 1993;11:191-212.
30. Brunner-Weinzierl MC, Hoff H, Burmester GR. Multiple functions for CD28 and cytotoxic T lymphocyte antigen-4 during different phases of T cell responses: Implications for arthritis and autoimmune diseases. Arthritis Res Ther 2004;6:45-54.
31. Zheng J, Yu X, Jiang L, Xiao M, Bai B, Lu J, Zhou Y. Association between the Cytotoxic T-lymphocyte antigen 4+ 49G > A polymorphism and cancer risk: a meta-analysis. BMC Cancer 2010;10:522.
32. Li D, Zhang Q, Xu F, Fu Z, Yuan W, Li D, Pang D. Association of CTLA-4 gene polymorphisms with sporadic breast cancer risk and clinical features in Han women of northeast China. Mol Cell Biochem 2012;364:283-90.
33. Li H, Fu ZK, Wang LH, Li DL, Wu N, Zhang J, Li DJ. Association of cytotoxic T lymphocyte antigen-4 gene polymorphisms with susceptibility to breast cancer. Xi Bao Yu Fen Zi Mian Yi Xue Za Zhi 2008;24:282-4. (in Chinese)
34. Bi Y, Wei L, Mao HT, Zhang L, Zuo WS. Expressions of Fas, CTLA-4 and RhoBTB2 genes in breast carcinoma and their relationship with clinicopathological factors. Zhonghua Zhong Liu Za Zhi 2008;30:749-53. (in Chinese)
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Farbod M, Shiryazdi SM, Harazi H, Nazari T, Sheikhha MH. Association between the cytotoxic T-lymphocyte antigen-4 polymorphisms and breast cancer risk and prognosis. J Cancer Metastasis Treat 2015;1:16-20. http://dx.doi.org/10.4103/2394-4722.153314
AMA Style
Farbod M, Shiryazdi SM, Harazi H, Nazari T, Sheikhha MH. Association between the cytotoxic T-lymphocyte antigen-4 polymorphisms and breast cancer risk and prognosis. Journal of Cancer Metastasis and Treatment. 2015; 1: 16-20. http://dx.doi.org/10.4103/2394-4722.153314
Chicago/Turabian Style
Farbod, Meraj, Seyed Mostafa Shiryazdi, Hamid Harazi, Tahereh Nazari, Mohammad Hasan Sheikhha. 2015. "Association between the cytotoxic T-lymphocyte antigen-4 polymorphisms and breast cancer risk and prognosis" Journal of Cancer Metastasis and Treatment. 1: 16-20. http://dx.doi.org/10.4103/2394-4722.153314
ACS Style
Farbod, M.; Shiryazdi SM.; Harazi H.; Nazari T.; Sheikhha MH. Association between the cytotoxic T-lymphocyte antigen-4 polymorphisms and breast cancer risk and prognosis. J. Cancer. Metastasis. Treat. 2015, 1, 16-20. http://dx.doi.org/10.4103/2394-4722.153314
About This Article
Copyright
Data & Comments
Data

 Cite This Article 2 clicks
Cite This Article 2 clicks











Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.